Convergence in τ
Last modified: 02-03-2026.
Go back to the Contents page.
Press Show to reveal the code chunks.
Let us set some global options for all code chunks in this document.
# Set seed for reproducibility
set.seed(1982)
# Set global options for all code chunks
knitr::opts_chunk$set(
# Disable messages printed by R code chunks
message = FALSE,
# Disable warnings printed by R code chunks
warning = FALSE,
# Show R code within code chunks in output
echo = TRUE,
# Include both R code and its results in output
include = TRUE,
# Evaluate R code chunks
eval = TRUE,
# Enable caching of R code chunks for faster rendering
cache = FALSE,
# Align figures in the center of the output
fig.align = "center",
# Enable retina display for high-resolution figures
retina = 2,
# Show errors in the output instead of stopping rendering
error = TRUE,
# Do not collapse code and output into a single block
collapse = FALSE
)
# Start the figure counter
fig_count <- 0
# Define the captioner function
captioner <- function(caption) {
fig_count <<- fig_count + 1
paste0("Figure ", fig_count, ": ", caption)
}To observe convergence in \(\tau\), we calibrate \(h\) and \(m\) to \(\tau\) as \(h \sim \tau^{1/\alpha}\) and \(m \sim \lceil\log_e^2(h)/(\pi^2(1-\alpha/2))\rceil\), respectively. Under this scaling, the total convergence rate with respect to the time step \(\tau\) is of order \(1\). That is, \(\|u-U_{h,m}^\tau\|_{L_2((0,T);L_2(\Gamma))} \leq C_\tau\tau\). This can be verified by estimating the slope \(S_\tau\) in the regression \(\log_{10} E = S_\tau\log_{10} \tau + \log_{10} C_\tau\), where we expect \(S_\tau\sim1\).
# remotes::install_github("davidbolin/rspde", ref = "devel")
# remotes::install_github("davidbolin/metricgraph", ref = "devel")
library(rSPDE)
library(MetricGraph)
library(grateful)
library(ggplot2)
library(reshape2)
library(plotly)
library(slackr)
source("keys.R")
slackr_setup(token = token) # token comes from keys.R## [1] "Successfully connected to Slack"capture.output(
knitr::purl(here::here("functionality.Rmd"), output = here::here("functionality.R")),
file = here::here("purl_log.txt")
)
source(here::here("functionality.R"))# Parameters
T_final <- 2
kappa <- 4
N_finite = 4 # choose even
adjusted_N_finite <- N_finite + N_finite/2 + 1
# Coefficients for u_0 and f
coeff_U_0 <- 50*(1:adjusted_N_finite)^-1
coeff_U_0[-5] <- 0
coeff_FF <- rep(0, adjusted_N_finite)
coeff_FF[7] <- 10
AAA = 1
OMEGA = pi
# Time step and mesh size
POWERS <- seq(6.125, 5.625, by = -0.1)
tau_vector <- 0.1 * 2^-POWERS
alpha_vector <- seq(1, 1.8, by = 0.2)# Overkill parameters
overkill_time_step <- 0.1 * 2^-14
overkill_h <- (0.1 * 2^-14)^(1/2)
# Finest time and space mesh
overkill_time_seq <- seq(0, T_final, length.out = ((T_final - 0) / overkill_time_step + 1))
overkill_graph <- gets.graph.tadpole(h = overkill_h)
# Compute the weights on the finest mesh
overkill_graph$compute_fem() # This is needed to compute the weights
overkill_C <- overkill_graph$mesh$C
m_values <- c()# Create a matrix to store the errors
errors_projected <- matrix(NA, nrow = length(tau_vector), ncol = length(alpha_vector))
for (j in 1:length(alpha_vector)) {
alpha <- alpha_vector[j] # from 0.5 to 2
beta <- alpha / 2
# Compute the eigenvalues and eigenfunctions on the finest mesh
overkill_eigen_params <- gets.eigen.params(N_finite = N_finite, kappa = kappa, alpha = alpha, graph = overkill_graph)
EIGENVAL_ALPHA <- overkill_eigen_params$EIGENVAL_ALPHA # Eigenvalues (they are independent of the meshes)
overkill_EIGENFUN <- overkill_eigen_params$EIGENFUN # Eigenfunctions on the finest mesh
# Compute the true solution on the finest mesh
overkill_U_true <- overkill_EIGENFUN %*%
outer(1:length(coeff_U_0),
1:length(overkill_time_seq),
function(i, j) (coeff_U_0[i] + coeff_FF[i] * G_sin(t = overkill_time_seq[j], A = AAA, lambda_j_alpha_half = EIGENVAL_ALPHA[i], omega = OMEGA)) * exp(-EIGENVAL_ALPHA[i] * overkill_time_seq[j]))
for (i in 1:length(tau_vector)) {
time_step <- tau_vector[i]
h <- time_step^(1/alpha)
m <- min(4, ceiling((log(h))^2 / (pi^2 * (1 - alpha / 2))))
#h <- largest_nested_h(h_star, h) # this makes the spatial mesh nested
m_values <- c(m_values, m)
graph <- gets.graph.tadpole(h = h)
graph$compute_fem()
G <- graph$mesh$G
C <- graph$mesh$C
L <- kappa^2*C + G
eigen_params <- gets.eigen.params(N_finite = N_finite, kappa = kappa, alpha = alpha, graph = graph)
EIGENFUN <- eigen_params$EIGENFUN
U_0 <- EIGENFUN %*% coeff_U_0 # Compute U_0 on the current mesh
Psi <- graph$fem_basis(overkill_graph$get_mesh_locations())
time_seq <- seq(0, T_final, length.out = ((T_final - 0) / time_step + 1))
my_op_frac <- my.fractional.operators.frac(L, beta, C, scale.factor = kappa^2, m = m, time_step)
INT_BASIS_EIGEN <- t(overkill_EIGENFUN) %*% overkill_C %*% Psi
# Compute matrix F with columns F^k
FF_approx <- t(INT_BASIS_EIGEN) %*%
outer(1:length(coeff_FF),
1:length(time_seq),
function(i, j) coeff_FF[i] * g_sin(r = time_seq[j], A = AAA, omega = OMEGA))
U_approx <- solve_fractional_evolution(my_op_frac, time_step, time_seq, val_at_0 = U_0, RHST = FF_approx)
projected_U_approx <- Psi %*% U_approx
projected_U_piecewise <- construct_piecewise_projection(projected_U_approx, time_seq, overkill_time_seq)
XX <- overkill_U_true - projected_U_piecewise
errors_projected[i,j] <- sqrt(as.double(overkill_time_step * sum(XX * (overkill_C %*% XX))))
slackr_msg(text = paste0("m =", m, ", alpha =", alpha, ", h =", h, ", time_step =", time_step), channel = "#research")
}
}
save(errors_projected, file = here::here("data_files/errors_projected_tau.RData"))observed_rates <- numeric(length(alpha_vector))
for (u in 1:length(alpha_vector)) {
observed_rates[u] <- coef(lm(log10(errors_projected[, u]) ~ log10(tau_vector)))[2]
}
theoretical_rates <- rep(1, length(alpha_vector))
p_tau <- error.convergence.plotter(x_axis_vector = tau_vector,
alpha_vector,
errors_projected,
theoretical_rates,
observed_rates,
line_equation_fun = loglog_line_equation,
fig_title = expression("Convergence in " * italic(tau)),
x_axis_label = expression(italic(tau)))
p_tau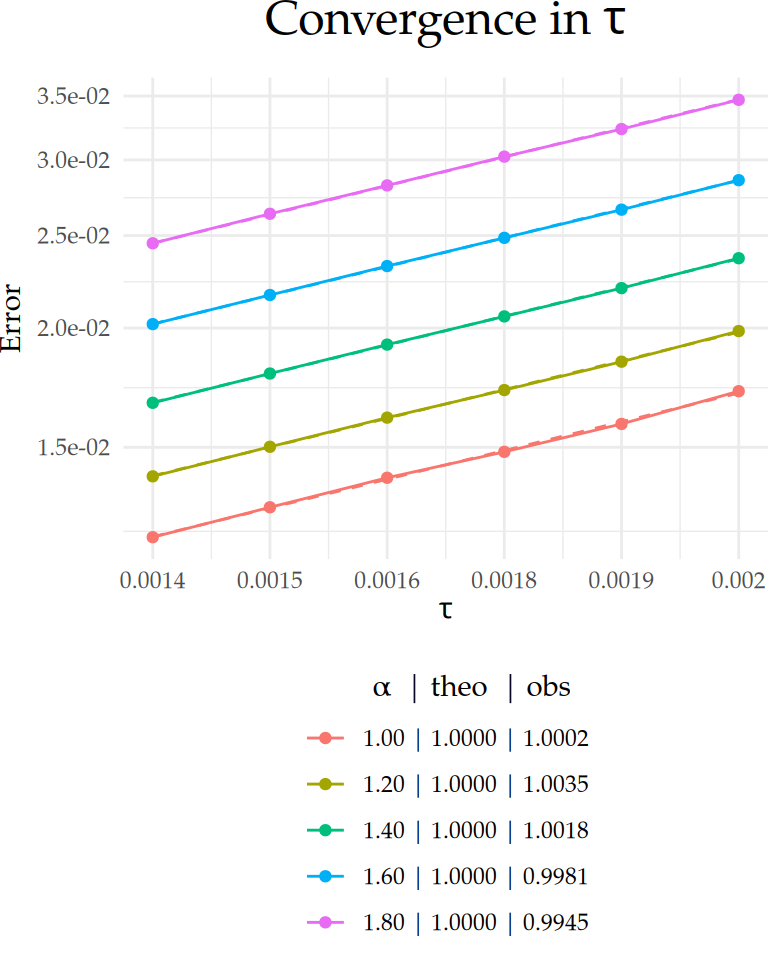
Figure 1: Comparison between theoretical and observed convergence behavior for the \(L_2(\Gamma\times(0,T))\)-error with respect to \(\tau\) on a \(\text{log}_{10}\)–\(\text{log}_{10}\) scale. Dashed lines indicate the theoretical rates, and solid lines represent the observed error curves. The legend below each plot shows the value of \(\alpha\) along with the corresponding theoretical (‘theo’), and observed (‘obs’) rates for each case.
ggsave(here::here("data_files/conv_rates_tau.png"), width = 4, height = 5, plot = p_tau, dpi = 300)
save(p_tau, file = here::here("data_files/p_tau.RData"))1 References
We used R version 4.5.2 (R Core Team 2025) and the following R packages: akima v. 0.6.3.6 (Akima and Gebhardt 2025), expm v. 1.0.0 (Maechler, Dutang, and Goulet 2024), fmesher v. 0.5.0 (Lindgren 2025), gsignal v. 0.3.7 (Van Boxtel, G.J.M., et al. 2021), here v. 1.0.1 (Müller 2020), htmltools v. 0.5.8.1 (Cheng et al. 2024), INLA v. 25.11.22 (Rue, Martino, and Chopin 2009; Lindgren, Rue, and Lindström 2011; Martins et al. 2013; Lindgren and Rue 2015; De Coninck et al. 2016; Rue et al. 2017; Verbosio et al. 2017; Bakka et al. 2018; Kourounis, Fuchs, and Schenk 2018), inlabru v. 2.13.0 (Yuan et al. 2017; Bachl et al. 2019), knitr v. 1.50 (Xie 2014, 2015, 2025a), Matrix v. 1.7.3 (Bates, Maechler, and Jagan 2025), MetricGraph v. 1.5.0.9000 (Bolin, Simas, and Wallin 2023a, 2023b, 2024, 2025; Bolin et al. 2024), neuralnet v. 1.44.2 (Fritsch, Guenther, and Wright 2019), orthopolynom v. 1.0.6.1 (Novomestky 2022), patchwork v. 1.3.1 (Pedersen 2025), pbmcapply v. 1.5.1 (Kuang, Kong, and Napolitano 2022), plotly v. 4.11.0 (Sievert 2020), posterdown v. 1.0 (Thorne 2019), pracma v. 2.4.4 (Borchers 2023), qrcode v. 0.3.0 (Onkelinx and Teh 2024), RColorBrewer v. 1.1.3 (Neuwirth 2022), RefManageR v. 1.4.0 (McLean 2014, 2017), renv v. 1.1.5 (Ushey and Wickham 2025), reshape2 v. 1.4.4 (Wickham 2007), reticulate v. 1.44.1 (Ushey, Allaire, and Tang 2025), rmarkdown v. 2.30 (Xie, Allaire, and Grolemund 2018; Xie, Dervieux, and Riederer 2020; Allaire et al. 2025), rSPDE v. 2.5.1.9000 (Bolin and Kirchner 2020; Bolin and Simas 2023; Bolin, Simas, and Xiong 2024), RSpectra v. 0.16.2 (Qiu and Mei 2024), scales v. 1.4.0 (Wickham, Pedersen, and Seidel 2025), slackr v. 3.4.0 (Kaye et al. 2025), tidyverse v. 2.0.0 (Wickham et al. 2019), viridisLite v. 0.4.2 (Garnier et al. 2023), xaringan v. 0.31 (Xie 2025b), xaringanExtra v. 0.8.0 (Aden-Buie and Warkentin 2024), xaringanthemer v. 0.4.4 (Aden-Buie 2025).