Go back to the Contents page.
Press Show to reveal the code chunks.
Go back to the About page.
This vignette processes half of the PeMS dataset, shows some
illustrations, and produces pems_repl1_data.RData,
which is a file that contains the graph with data and a covariate
defined on the mesh. To see how the data is modeled, go to pems_repl2.html.
Let us set some global options for all code chunks in this
document.
# Create a clipboard button on the rendered HTML page
source(here::here("clipboard.R")); clipboard
# Set seed for reproducibility
set.seed(1982)
# Set global options for all code chunks
knitr::opts_chunk$set(
# Disable messages printed by R code chunks
message = FALSE,
# Disable warnings printed by R code chunks
warning = FALSE,
# Show R code within code chunks in output
echo = TRUE,
# Include both R code and its results in output
include = TRUE,
# Evaluate R code chunks
eval = FALSE,
# Enable caching of R code chunks for faster rendering
cache = FALSE,
# Align figures in the center of the output
fig.align = "center",
# Enable retina display for high-resolution figures
retina = 2,
# Show errors in the output instead of stopping rendering
error = TRUE,
# Do not collapse code and output into a single block
collapse = FALSE
)
# Start the figure counter
fig_count <- 0
# Define the captioner function
captioner <- function(caption) {
fig_count <<- fig_count + 1
paste0("Figure ", fig_count, ": ", caption)
}
# Define the function to truncate a number to two decimal places
truncate_to_two <- function(x) {
truncated <- floor(x * 100) / 100
sprintf("%.2f", truncated)
}
Below we load the necessary libraries.
library(INLA)
library(inlabru)
library(rSPDE)
library(MetricGraph)
library(dplyr)
library(plotly)
library(scales)
library(patchwork)
library(ggplot2)
library(cowplot)
library(ggpubr) #annotate_figure()
library(grid) #textGrob()
library(ggmap)
library(viridis)
library(OpenStreetMap)
library(tidyr)
library(sf)
library(here)
library(rmarkdown)
library(grateful) # Cite all loaded packages
library(slackr)
source("keys.R")
slackr_setup(token = token) # token comes from keys.R
## [1] "Successfully connected to Slack"
capture.output(
knitr::purl(here::here("functionality1.Rmd"), output = here::here("functionality1.R")),
file = here::here("old/purl_log.txt")
)
source(here::here("functionality1.R"))
Below we define function gets_summary_parameters() to
extract the summary of the parameters of the model.
gets_summary_parameters <- function(fit, model) {
param_spde <- summary(rspde.result(fit, "field", model, parameterization = "spde"))
param_matern <- summary(rspde.result(fit, "field", model, parameterization = "matern"))
param_fixed <- fit$summary.fixed[,1:6]
marginal.posterior.sigma_e = inla.tmarginal(
fun = function(x) exp(-x/2),
marginal = fit[["internal.marginals.hyperpar"]][["Log precision for the Gaussian observations"]])
quant.sigma_e <- capture.output({result_tmp <- inla.zmarginal(marginal.posterior.sigma_e)}, file = "/dev/null")
quant.sigma_e <- result_tmp
statistics.sigma_e <- unlist(quant.sigma_e)[c(1,2,3,5,7)]
mode.sigma_e <- inla.mmarginal(marginal.posterior.sigma_e)
allparams <- rbind(param_fixed, param_spde, param_matern, c(statistics.sigma_e, mode.sigma_e))
rownames(allparams)[nrow(allparams)] <- "sigma_e"
return(allparams)
}
Folder data.pems was downloaded from this GitHub
repository.
Lines <- read_sf(here("data_files/data.pems/lines.shp"))
lines <- as_Spatial(Lines)
EtV <- read.csv(here("data_files/data.pems/E.csv"), header = T, row.names = NULL)
PtE <- read.csv(here("data_files/data.pems/PtE.csv"), header = T, row.names = NULL)
PtE[,1] <- PtE[,1] + 1
Y <- read.csv(here("data_files/data.pems/Y.csv"), header = T, row.names = NULL)
Y <- as.matrix(Y[,-1])
edge_length_m <- EtV[,4]
PtE[,2] = PtE[,2]/edge_length_m[PtE[,1]]
Matrix Y has dimension \(26\times325\). Each row of Y
corresponds to a replicate (\(26\)
replicates in total) and each column corresponds to a location (\(325\) locations in total) on the
network.
We remove columns in Y, those with the same value in the
first 13 replicates. We also remove the corresponding rows in
PtE.
all_same <- function(col) {
length(unique(col[1:13])) == 1
}
cols_to_keep <- apply(Y, 2, function(col) !all_same(col))
Y <- Y[, cols_to_keep]
PtE <- PtE[cols_to_keep,]
Half of the replicates (Y_for_summary) are used to
compute Y_logstd and Y_mean, and the other
half (Y again ) are used to fit a replicate model in pems_repl2.html.
sampled_numbers <- 1:13
not_sampled_numbers <- 14:26
Y_for_summary <- Y[sampled_numbers,]
Y_logstd <- apply(Y_for_summary, 2, sd) |> as.vector() |> log()
Y_mean <- apply(Y_for_summary, 2, mean) |> as.vector()
Y <- Y[not_sampled_numbers,]
save(Y_mean, file = here::here("data_files/Y_mean.RData"))
Observe that Y_logstd corresponds to \(\log\) of the standard deviation at each
location.
Below we build the graph using the list element edges
object from pems_repl.
# Build the graph
graph <- metric_graph$new(edges = pems_repl$edges, longlat = TRUE)
# Add the observations
graph$add_observations(data = data.frame(y = Y_logstd,
edge_number = PtE[,1],
distance_on_edge = PtE[,2]),
edge_number = "edge_number",
distance_on_edge = "distance_on_edge",
data_coords = "PtE",
normalized = TRUE,
clear_obs = TRUE)
# Build the mesh
graph$build_mesh(h = 0.05)
Below we fit \(\log(\mathrm{sd}(s_i))\sim
N(\beta_0+u(s_i), \sigma_e^2)\).
# Build the model
rspde_model_stat_logstd <- rspde.metric_graph(graph,
parameterization = "spde")
# Prepare the data for fitting
data_rspde_bru_stat <- graph_data_rspde(rspde_model_stat_logstd, bru = TRUE)
# Define the component
cmp_stat <- y ~ -1 +
Intercept(1) +
field(cbind(.edge_number, .distance_on_edge), model = rspde_model_stat_logstd)
# Fit the model
rspde_fit_stat_logstd <-
bru(cmp_stat,
data = data_rspde_bru_stat[["data"]],
family = "gaussian",
options = list(verbose = FALSE)
)
save(rspde_fit_stat_logstd, rspde_model_stat_logstd, file = here::here("data_files/rspde_fit_stat_and_model_logstd.RData"))
load(here::here("data_files/rspde_fit_stat_and_model_logstd.RData"))
# Summarize the results
summary(rspde_fit_stat_logstd)
## inlabru version: 2.13.0
## INLA version: 25.11.22
## Components:
## Intercept: main = linear(1), group = exchangeable(1L), replicate = iid(1L), NULL
## field: main = cgeneric(cbind(.edge_number, .distance_on_edge)), group = exchangeable(1L), replicate = iid(1L), NULL
## Observation models:
## Family: 'gaussian'
## Tag: <No tag>
## Data class: 'metric_graph_data', 'data.frame'
## Response class: 'numeric'
## Predictor: y ~ .
## Additive/Linear: TRUE/TRUE
## Used components: effects[Intercept, field], latent[]
## Time used:
## Pre = 0.549, Running = 3.27, Post = 1.19, Total = 5.01
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## Intercept 1.756 0.133 1.491 1.756 2.02 1.756 0
##
## Random effects:
## Name Model
## field CGeneric
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 4.600 0.504 3.685 4.574
## Theta1 for field 0.261 0.193 -0.120 0.261
## Theta2 for field -0.647 0.224 -1.084 -0.649
## Theta3 for field 0.886 0.688 -0.432 0.873
## 0.975quant mode
## Precision for the Gaussian observations 5.667 4.523
## Theta1 for field 0.642 0.261
## Theta2 for field -0.201 -0.655
## Theta3 for field 2.278 0.819
##
## Marginal log-Likelihood: -327.36
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')
gets_summary_parameters(rspde_fit_stat_logstd, rspde_model_stat_logstd)
We now compute the kriging predictor \(E(u(s_i)|\log(\mathrm{sd}(s_i)))\) (denoted
as cov below) and standardize it.
# Prediction locations
data_prd_list_mesh <- graph$get_mesh_locations(bru = TRUE)
# Compute the kriging predictor
y_pred <- predict(rspde_fit_stat_logstd, newdata = data_prd_list_mesh, ~Intercept + field)
cov_almost <- y_pred$mean
# Standardize the kriging predictor of the log standard deviation
cov <- (cov_almost - mean(cov_almost))/sd(cov_almost)
Below we plot the results. We first generate two maps, one for zoom
level 12 and the other for zoom level 13.
# Extract the Euclidean coordinates of the mesh points
xypoints <- graph$mesh$V
# Extract the range of the coordinates
x_left <- range(xypoints[,1])[1]
x_right <- range(xypoints[,1])[2]
y_bottom <- range(xypoints[,2])[1]
y_top <- range(xypoints[,2])[2]
# Set the style of the map
style_vector <- c(c(feature = "road", element = "geometry", visibility = "on"),
c("&style=", feature = "poi", element = "labels", visibility = "off"),
c("&style=", feature = "road", element = "labels", visibility = "off"),
c("&style=", feature = "administrative", element = "labels", visibility = "off"), #name of places
c("&style=", feature = "transit", element = "labels", visibility = "off"),
c("&style=", feature = "landscape", element = "geometry", visibility = "on"),
c("&style=", feature = "water", element = "geometry", visibility = "on"))
register_stadiamaps(Sys.getenv("GGMAP_STADIAMAPS_API_KEY"))
# Obtain the maps
p12 <- ggmap(get_stadiamap(bbox = c(left = x_left, bottom = y_bottom, right = x_right, top = y_top),
zoom = 12,
maptype = "stamen_toner_lite",
scale =2,
style = style_vector,
color = "color")) + xlab(NULL) + ylab(NULL)
p13 <- ggmap(get_stadiamap(bbox = c(left = x_left, bottom = y_bottom, right = x_right, top = y_top),
zoom = 13,
maptype = "stamen_toner_lite",
scale =2,
style = style_vector,
color = "color")) + xlab(NULL) + ylab(NULL)
save(p12, p13, file = here::here("data_files/maps_zoom12and13from_stadia.RData"))
load(here::here("data_files/maps_zoom12and13from_stadia.RData"))
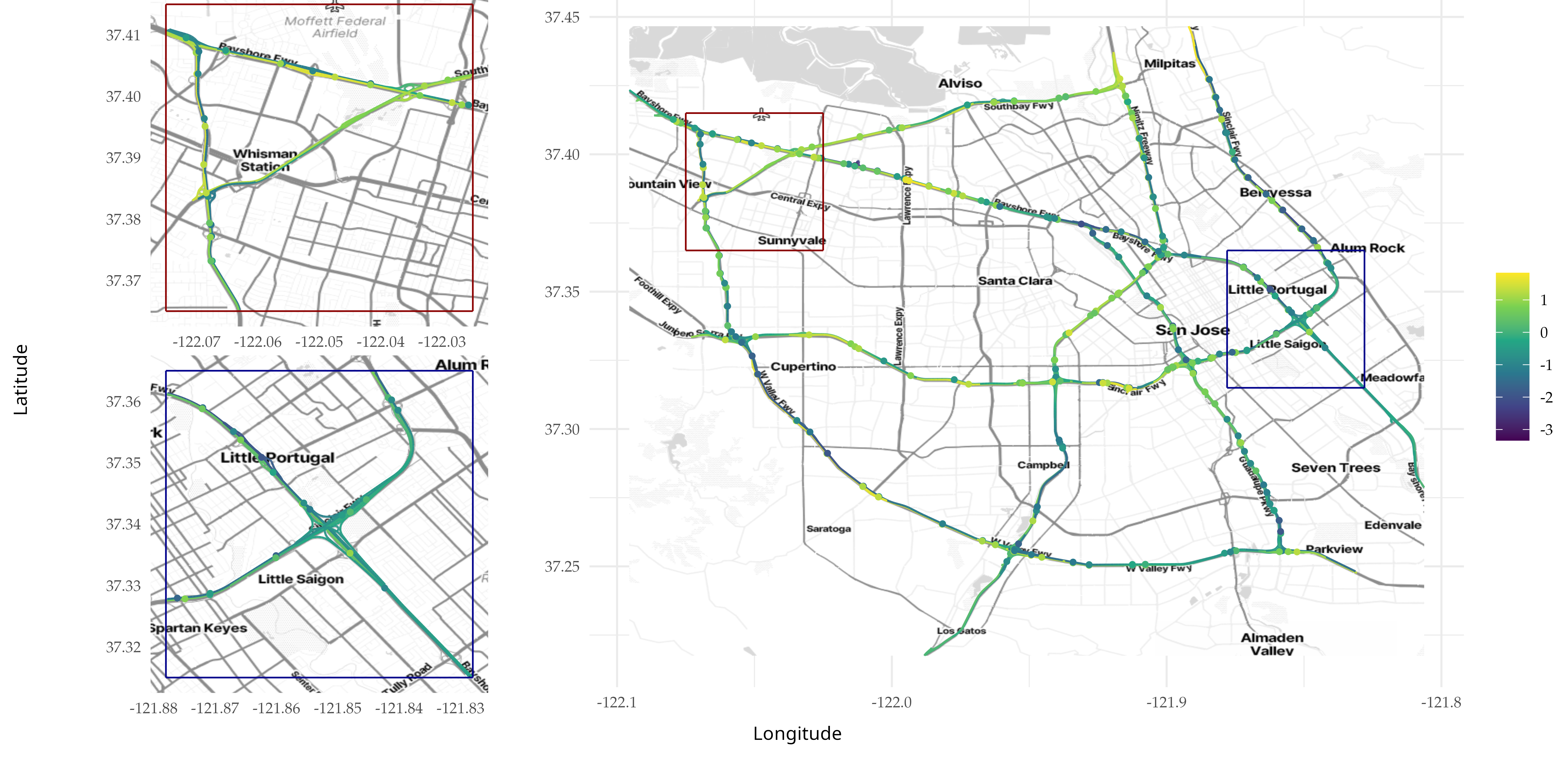
Below we plot the field on top of the map. We also add the point
values of the standard deviation of the log standard deviation to the
graph so we can plot it on top of the map and the field.
# Define the colors for the windows
lower_color <- "darkred"
upper_color <- "darkblue"
# Define coordinates for small windows
coordx_lwr1 <- -121.878
coordx_upr1 <- -121.828
coordy_lwr1 <- 37.315
coordy_upr1 <- 37.365
coordx_lwr2<- -122.075
coordx_upr2 <- -122.025
coordy_lwr2 <- 37.365
coordy_upr2 <- 37.415
# Plot the field on top of the map
f12 <- graph$plot_function(X = cov,
vertex_size = 0,
p = p12,
edge_width = 0.5) +
theme_minimal() +
theme(text = element_text(family = "Palatino"),
axis.text = element_text(size = 8),
legend.text = element_text(size = 8),
plot.margin = unit(-0.4*c(1,0,1,1), "cm")
) +
labs(color = "", x = "", y = "") +
xlim(x_left, x_right) +
ylim(y_bottom, y_top)
# Plot the field on top of the map
f13 <- graph$plot_function(X = cov,
vertex_size = 0,
p = p13,
edge_width = 0.5) +
theme_minimal() +
theme(text = element_text(family = "Palatino"),
axis.text = element_text(size = 8),
legend.text = element_text(size = 8),
plot.margin = unit(-0.4*c(1,0,1,1), "cm")
) +
labs(color = "", x = "", y = "") +
xlim(x_left, x_right) +
ylim(y_bottom, y_top)
# We add the point values of standard deviation of the log standard deviation to the graph so we can plot it
standardized_Y_logstd <- (Y_logstd - mean(Y_logstd))/sd(Y_logstd)
graph$add_observations(data = data.frame(y = standardized_Y_logstd,
edge_number = PtE[,1],
distance_on_edge = PtE[,2]),
edge_number = "edge_number",
distance_on_edge = "distance_on_edge",
data_coords = "PtE",
normalized = TRUE,
clear_obs = TRUE)
g12 <- graph$plot(data = "y", vertex_size = 0, p = f12, edge_width = 0, data_size = 1) +
labs(color = "", x = "", y = "") +
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_lwr1,
linewidth = 0.4, color = upper_color) + # Bottom line
annotate("segment", x = coordx_lwr1, y = coordy_upr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Top line
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_lwr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Left line
annotate("segment", x = coordx_upr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Right line
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_lwr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_upr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_lwr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_upr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color)
g13 <- graph$plot(data = "y", vertex_size = 0, p = f13, edge_width = 0, data_size = 1) +
labs(color = "", x = "", y = "") +
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_lwr1,
linewidth = 0.4, color = upper_color) + # Bottom line
annotate("segment", x = coordx_lwr1, y = coordy_upr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Top line
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_lwr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Left line
annotate("segment", x = coordx_upr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Right line
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_lwr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_upr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_lwr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_upr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color)
r1 <- g13 + xlim(coordx_lwr1, coordx_upr1) +
ylim(coordy_lwr1, coordy_upr1) +
theme(legend.position = "none",
plot.margin = unit(-0.2*c(1,1,1,1), "cm"))
r2 <- g13 + xlim(coordx_lwr2, coordx_upr2) +
ylim(coordy_lwr2, coordy_upr2) +
theme(legend.position = "none",
plot.margin = unit(-0.2*c(1,1,1,1), "cm"))
# Arrange p2 and p3 horizontally
left_col <- plot_grid(r2, r1, labels = NULL, ncol = 1, nrow = 2, rel_heights = c(1,1))
# Combine the top row with p1 in a grid
combined_plot <- plot_grid(left_col, g12, labels = NULL, ncol = 2, rel_widths = c(1,2))
final_plot <- annotate_figure(combined_plot, left = textGrob("Latitude", rot = 90, vjust = 1, gp = gpar(cex = 0.8)),
bottom = textGrob("Longitude", vjust = -0.5, gp = gpar(cex = 0.8)))
ggsave(here("data_files/stand_log_sigma_3.png"), width = 11.2, height = 5.43, plot = final_plot, dpi = 500)
save(final_plot, file = here("data_files/stand_log_sigma_3.RData"))
# Print the combined plot
knitr::include_graphics(here::here("data_files/stand_log_sigma_3.png"))
We repeat the same process but now for Y_mean. We first
add the observations to the graph.
graph$add_observations(data = data.frame(y = Y_mean,
edge_number = PtE[,1],
distance_on_edge = PtE[,2]),
edge_number = "edge_number",
distance_on_edge = "distance_on_edge",
data_coords = "PtE",
normalized = TRUE,
clear_obs = TRUE)
We fit the model \(\mathrm{mean}(s_i)\sim
N(\beta_0+u(s_i), \sigma_e^2)\).
# Build the model
rspde_model_stat_mean <- rspde.metric_graph(graph,
parameterization = "spde")
# Prepare the data for fitting
data_rspde_bru_stat <- graph_data_rspde(rspde_model_stat_mean,
bru = TRUE)
# Define the component
cmp_stat <- y ~ -1 +
Intercept(1) +
field(cbind(.edge_number, .distance_on_edge), model = rspde_model_stat_mean)
# Fit the model
rspde_fit_stat_mean <-
bru(cmp_stat,
data = data_rspde_bru_stat[["data"]],
family = "gaussian",
options = list(verbose = FALSE)
)
save(rspde_fit_stat_mean, rspde_model_stat_mean, file = here::here("data_files/rspde_fit_stat_and_model_mean.RData"))
load(here::here("data_files/rspde_fit_stat_and_model_mean.RData"))
# Summarize the results
summary(rspde_fit_stat_mean)
## inlabru version: 2.13.0
## INLA version: 25.11.22
## Components:
## Intercept: main = linear(1), group = exchangeable(1L), replicate = iid(1L), NULL
## field: main = cgeneric(cbind(.edge_number, .distance_on_edge)), group = exchangeable(1L), replicate = iid(1L), NULL
## Observation models:
## Family: 'gaussian'
## Tag: <No tag>
## Data class: 'metric_graph_data', 'data.frame'
## Response class: 'numeric'
## Predictor: y ~ .
## Additive/Linear: TRUE/TRUE
## Used components: effects[Intercept, field], latent[]
## Time used:
## Pre = 0.538, Running = 8.84, Post = 0.626, Total = 10
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## Intercept 50.758 2.945 44.851 50.778 56.538 50.776 0
##
## Random effects:
## Name Model
## field CGeneric
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 0.020 0.002 0.016 0.019
## Theta1 for field -2.326 0.133 -2.603 -2.321
## Theta2 for field -0.753 0.261 -1.224 -0.766
## Theta3 for field 1.209 0.542 0.256 1.175
## 0.975quant mode
## Precision for the Gaussian observations 0.024 0.019
## Theta1 for field -2.081 -2.297
## Theta2 for field -0.199 -0.830
## Theta3 for field 2.376 1.010
##
## Marginal log-Likelihood: -1193.09
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')
gets_summary_parameters(rspde_fit_stat_mean, rspde_model_stat_mean)
Below we compute the predictions for the mesh and the data
location.
mean_on_mesh_pred <- predict(rspde_fit_stat_mean, newdata = data_prd_list_mesh, ~Intercept + field)
cov_for_mean_to_plot <- mean_on_mesh_pred$mean
mean_on_loc_pred <- predict(rspde_fit_stat_mean, newdata = rename(PtE, .edge_number = E, .distance_on_edge = length), ~Intercept + field)
cov_for_mean <- mean_on_loc_pred$mean
save(cov_for_mean_to_plot, cov_for_mean, file = here::here("data_files/cov_for_mean_to_plot_and_cov_for_mean.RData"))
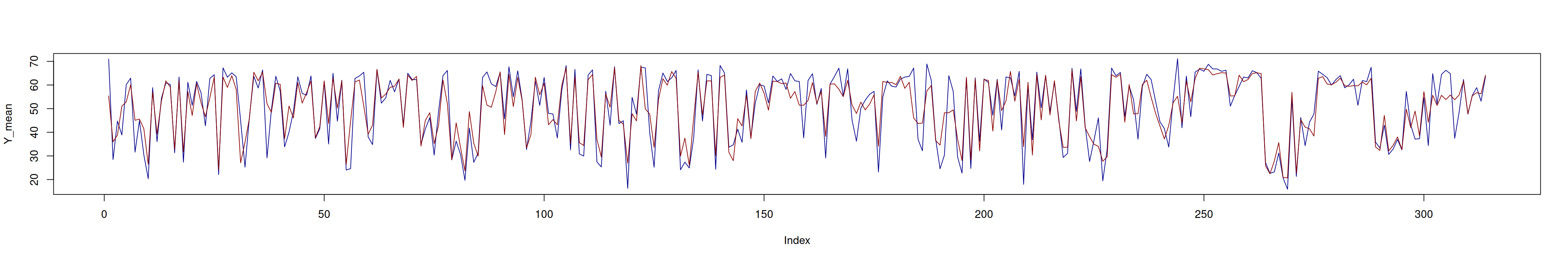
Below we check that the predicted mean is close to the observed
mean.
load(here::here("data_files/Y_mean.RData"))
load(here::here("data_files/cov_for_mean_to_plot_and_cov_for_mean.RData"))
plot(Y_mean, type = "l", col = "darkblue")
lines(cov_for_mean, col = "darkred")
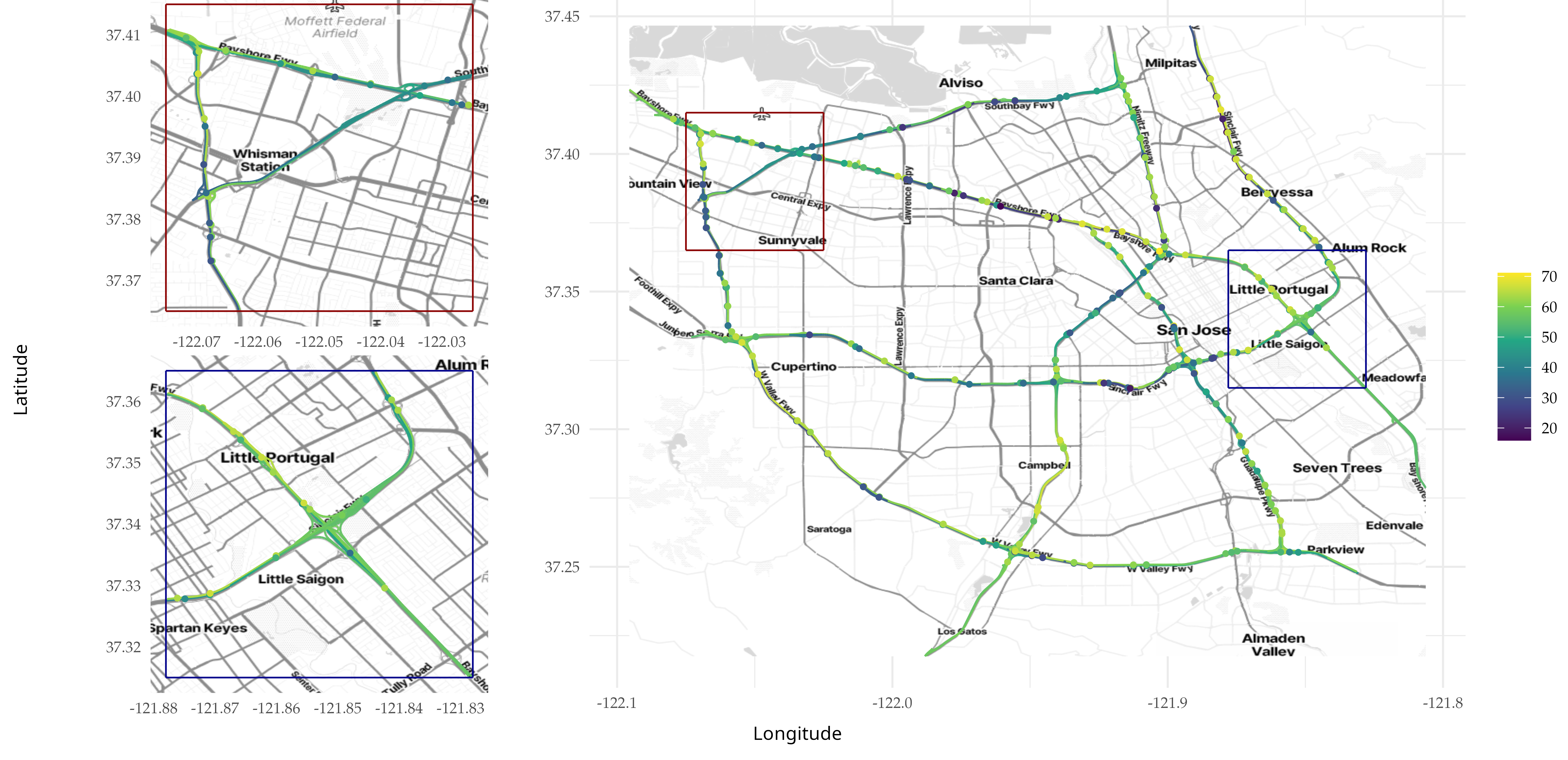
Below we plot the results as we did before.
# Plot the field on top of the map
f12 <- graph$plot_function(X = cov_for_mean_to_plot,
vertex_size = 0,
p = p12,
edge_width = 0.5) +
theme_minimal() +
theme(text = element_text(family = "Palatino"),
axis.text = element_text(size = 8),
legend.text = element_text(size = 8),
plot.margin = unit(-0.4*c(1,0,1,1), "cm")
) +
labs(color = "", x = "", y = "") +
xlim(x_left, x_right) +
ylim(y_bottom, y_top)
# Plot the field on top of the map
f13 <- graph$plot_function(X = cov_for_mean_to_plot,
vertex_size = 0,
p = p13,
edge_width = 0.5) +
theme_minimal() +
theme(text = element_text(family = "Palatino"),
axis.text = element_text(size = 8),
legend.text = element_text(size = 8),
plot.margin = unit(-0.4*c(1,0,1,1), "cm")
) +
labs(color = "", x = "", y = "") +
xlim(x_left, x_right) +
ylim(y_bottom, y_top)
g12 <- graph$plot(data = "y", vertex_size = 0, p = f12, edge_width = 0, data_size = 1) +
labs(color = "", x = "", y = "") +
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_lwr1,
linewidth = 0.4, color = upper_color) + # Bottom line
annotate("segment", x = coordx_lwr1, y = coordy_upr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Top line
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_lwr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Left line
annotate("segment", x = coordx_upr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Right line
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_lwr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_upr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_lwr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_upr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color)
g13 <- graph$plot(data = "y", vertex_size = 0, p = f13, edge_width = 0, data_size = 1) +
labs(color = "", x = "", y = "") +
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_lwr1,
linewidth = 0.4, color = upper_color) + # Bottom line
annotate("segment", x = coordx_lwr1, y = coordy_upr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Top line
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_lwr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Left line
annotate("segment", x = coordx_upr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Right line
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_lwr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_upr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_lwr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_upr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color)
r1 <- g13 + xlim(coordx_lwr1, coordx_upr1) +
ylim(coordy_lwr1, coordy_upr1) +
theme(legend.position = "none",
plot.margin = unit(-0.2*c(1,1,1,1), "cm"))
r2 <- g13 + xlim(coordx_lwr2, coordx_upr2) +
ylim(coordy_lwr2, coordy_upr2) +
theme(legend.position = "none",
plot.margin = unit(-0.2*c(1,1,1,1), "cm"))
# Arrange p2 and p3 horizontally
left_col <- plot_grid(r2, r1, labels = NULL, ncol = 1, nrow = 2, rel_heights = c(1,1))
# Combine the top row with p1 in a grid
combined_plot <- plot_grid(left_col, g12, labels = NULL, ncol = 2, rel_widths = c(1,2))
final_plot <- annotate_figure(combined_plot, left = textGrob("Latitude", rot = 90, vjust = 1, gp = gpar(cex = 0.8)),
bottom = textGrob("Longitude", vjust = -0.5, gp = gpar(cex = 0.8)))
ggsave(here("data_files/mean_better_3.png"), width = 11.2, height = 5.43, plot = final_plot, dpi = 500)
knitr::include_graphics(here::here("data_files/mean_better_3.png"))
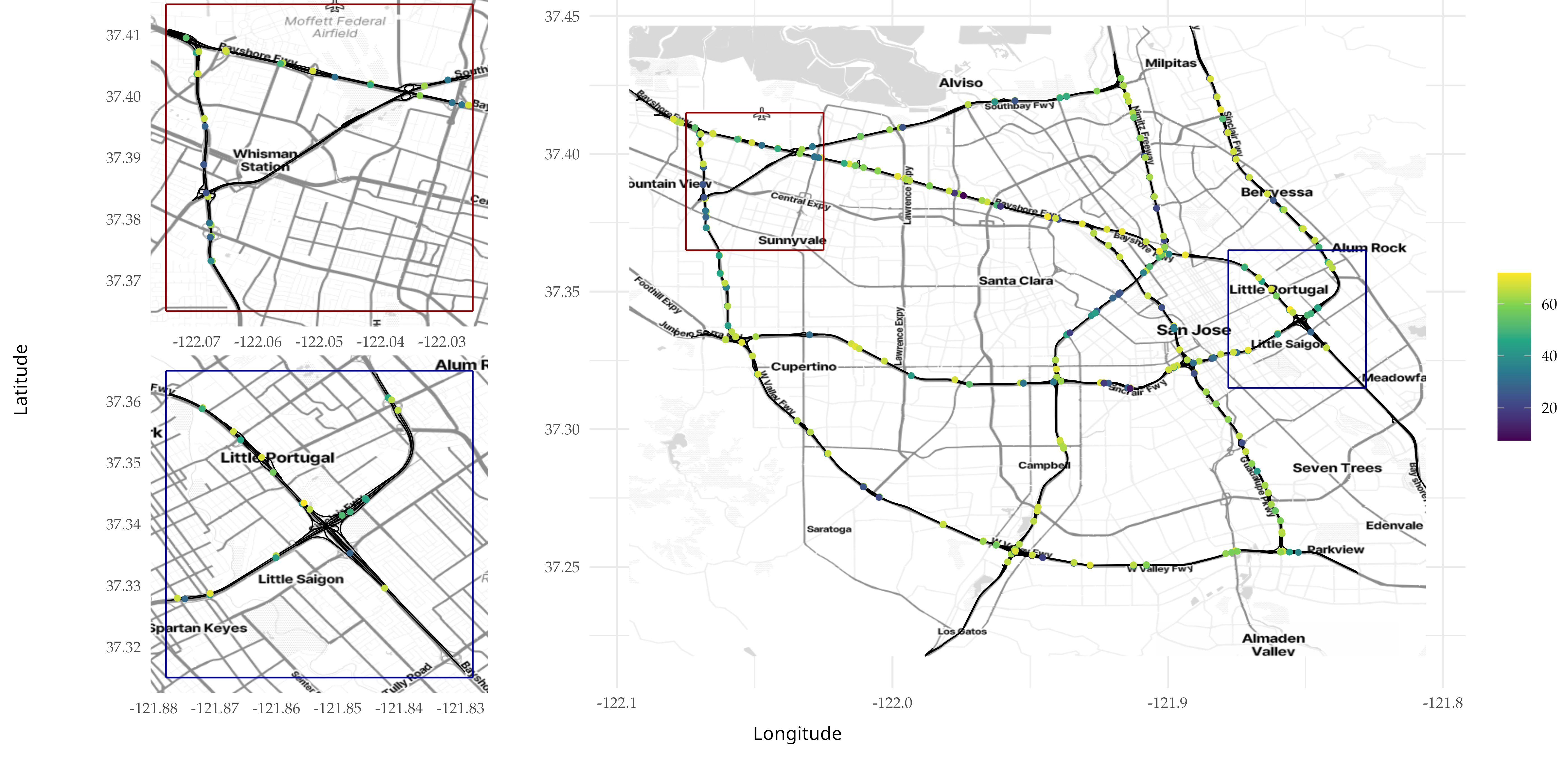
We finally add the remaining half of the replicates to the graph and
plot one of them.
df_rep <- lapply(1:nrow(Y), function(i){data.frame(y = Y[i,],
mean_value = cov_for_mean,
edge_number = PtE[,1],
distance_on_edge = PtE[,2],
repl = i)})
df_rep <- do.call(rbind, df_rep)
graph$add_observations(data = df_rep,
edge_number = "edge_number",
distance_on_edge = "distance_on_edge",
data_coords = "PtE",
normalized = TRUE,
clear_obs = TRUE,
group = "repl")
Below we plot replicate 14.
g12 <- graph$plot(data = "y",
p = p12,
group = 1,
vertex_size = 0,
data_size = 1) +
theme_minimal() +
theme(text = element_text(family = "Palatino"),
axis.text = element_text(size = 8),
legend.text = element_text(size = 8),
plot.margin = unit(-0.4*c(1,0,1,1), "cm")) +
labs(color = "", x = "", y = "") +
xlim(x_left, x_right) +
ylim(y_bottom, y_top) +
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_lwr1,
linewidth = 0.4, color = upper_color) + # Bottom line
annotate("segment", x = coordx_lwr1, y = coordy_upr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Top line
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_lwr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Left line
annotate("segment", x = coordx_upr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Right line
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_lwr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_upr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_lwr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_upr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color)
g13 <- graph$plot(data = "y",
p = p13,
group = 1,
vertex_size = 0,
data_size = 1) +
theme_minimal() +
theme(text = element_text(family = "Palatino"),
axis.text = element_text(size = 8),
legend.text = element_text(size = 8),
plot.margin = unit(-0.4*c(1,0,1,1), "cm")) +
labs(color = "speed", x = "", y = "") +
xlim(x_left, x_right) +
ylim(y_bottom, y_top) +
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_lwr1,
linewidth = 0.4, color = upper_color) + # Bottom line
annotate("segment", x = coordx_lwr1, y = coordy_upr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Top line
annotate("segment", x = coordx_lwr1, y = coordy_lwr1, xend = coordx_lwr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Left line
annotate("segment", x = coordx_upr1, y = coordy_lwr1, xend = coordx_upr1, yend = coordy_upr1,
linewidth = 0.4, color = upper_color) + # Right line
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_lwr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_upr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_lwr2, y = coordy_lwr2, xend = coordx_lwr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color) +
annotate("segment", x = coordx_upr2, y = coordy_lwr2, xend = coordx_upr2, yend = coordy_upr2,
linewidth = 0.4, color = lower_color)
r1 <- g13 + xlim(coordx_lwr1, coordx_upr1) +
ylim(coordy_lwr1, coordy_upr1) +
theme(legend.position = "none",
plot.margin = unit(-0.2*c(1,1,1,1), "cm"))
r2 <- g13 + xlim(coordx_lwr2, coordx_upr2) +
ylim(coordy_lwr2, coordy_upr2) +
theme(legend.position = "none",
plot.margin = unit(-0.2*c(1,1,1,1), "cm"))
# Arrange p2 and p3 horizontally
left_col <- plot_grid(r2, r1, labels = NULL, ncol = 1, nrow = 2, rel_heights = c(1,1))
# Combine the top row with p1 in a grid
combined_plot <- plot_grid(left_col, g12, labels = NULL, ncol = 2, rel_widths = c(1,2))
final_plot <- annotate_figure(combined_plot, left = textGrob("Latitude", rot = 90, vjust = 1, gp = gpar(cex = 0.8)),
bottom = textGrob("Longitude", vjust = -0.5, gp = gpar(cex = 0.8)))
ggsave(here("data_files/replicate14_3.png"), width = 11.2, height = 5.43, plot = final_plot, dpi = 500)
knitr::include_graphics(here::here("data_files/replicate14_3.png"))
save(graph, cov, cov_for_mean_to_plot, data_prd_list_mesh, file = here("data_files/pems_repl1_data.RData"))
References
grateful::cite_packages(output = "paragraph", out.dir = ".")
We used R version 4.5.2 (R Core Team
2025) and the following R packages: cowplot v. 1.2.0 (Wilke 2025), ggmap v. 4.0.2 (Kahle and Wickham 2013), ggpubr v. 0.6.3 (Kassambara 2026), ggtext v. 0.1.2 (Wilke and Wiernik 2022), glue v. 1.8.0 (Hester and Bryan 2024), here v. 1.0.1 (Müller 2020), htmltools v. 0.5.8.1 (Cheng et al. 2024), INLA v. 25.11.22 (Rue, Martino, and Chopin 2009; Lindgren, Rue, and
Lindström 2011; Martins et al. 2013; Lindgren and Rue 2015; De Coninck
et al. 2016; Rue et al. 2017; Verbosio et al. 2017; Bakka et al. 2018;
Kourounis, Fuchs, and Schenk 2018), inlabru v. 2.13.0 (Yuan et al. 2017; Bachl et al. 2019), knitr v.
1.50 (Xie 2014, 2015, 2025), latex2exp v.
0.9.8 (Meschiari 2026), Matrix v. 1.7.3
(Bates, Maechler, and Jagan 2025),
MetricGraph v. 1.5.0.9000 (Bolin, Simas, and
Wallin 2023a, 2023b, 2024, 2025; Bolin et al. 2024),
OpenStreetMap v. 0.4.1 (Fellows and Stotz
2025), patchwork v. 1.3.1 (Pedersen
2025), plotly v. 4.11.0 (Sievert
2020), plotrix v. 3.8.14 (J 2006),
renv v. 1.1.7 (Ushey and Wickham 2026),
reshape2 v. 1.4.4 (Wickham 2007),
reticulate v. 1.44.1 (Ushey, Allaire, and Tang
2025), rmarkdown v. 2.30 (Xie, Allaire,
and Grolemund 2018; Xie, Dervieux, and Riederer 2020; Allaire et al.
2025), rSPDE v. 2.5.2.9000 (Bolin and
Kirchner 2020; Bolin and Simas 2023; Bolin, Simas, and Xiong
2024), scales v. 1.4.0 (Wickham, Pedersen,
and Seidel 2025), sf v. 1.1.0 (E. Pebesma
2018; E. Pebesma and Bivand 2023), slackr v. 3.4.0 (Kaye et al. 2025), sp v. 2.2.1 (E. J. Pebesma and Bivand 2005; Bivand, Pebesma, and
Gomez-Rubio 2013), tidyverse v. 2.0.0 (Wickham et al. 2019), tikzDevice v. 0.12.6
(Sharpsteen and Bracken 2023), viridis v.
0.6.5 (Garnier et al. 2024), xaringanExtra
v. 0.8.0 (Aden-Buie and Warkentin
2024).
Allaire, JJ, Yihui Xie, Christophe Dervieux, Jonathan McPherson, Javier
Luraschi, Kevin Ushey, Aron Atkins, et al. 2025.
rmarkdown: Dynamic Documents for r.
https://github.com/rstudio/rmarkdown.
Bachl, Fabian E., Finn Lindgren, David L. Borchers, and Janine B.
Illian. 2019.
“inlabru: An
R Package for Bayesian Spatial Modelling from
Ecological Survey Data.” Methods in Ecology and
Evolution 10: 760–66.
https://doi.org/10.1111/2041-210X.13168.
Bakka, Haakon, Håvard Rue, Geir-Arne Fuglstad, Andrea I. Riebler, David
Bolin, Janine Illian, Elias Krainski, Daniel P. Simpson, and Finn K.
Lindgren. 2018.
“Spatial Modelling with INLA:
A Review.” WIRES (Invited Extended Review)
xx (Feb): xx–.
http://arxiv.org/abs/1802.06350.
Bates, Douglas, Martin Maechler, and Mikael Jagan. 2025.
Matrix: Sparse and Dense Matrix Classes and
Methods.
https://doi.org/10.32614/CRAN.package.Matrix.
Bivand, Roger S., Edzer Pebesma, and Virgilio Gomez-Rubio. 2013.
Applied Spatial Data Analysis with R, Second
Edition. Springer, NY.
https://asdar-book.org/.
Bolin, David, and Kristin Kirchner. 2020.
“The Rational
SPDE Approach for Gaussian Random Fields with
General Smoothness.” Journal of Computational and Graphical
Statistics 29 (2): 274–85.
https://doi.org/10.1080/10618600.2019.1665537.
Bolin, David, Mihály Kovács, Vivek Kumar, and Alexandre B. Simas. 2024.
“Regularity and Numerical Approximation of Fractional Elliptic
Differential Equations on Compact Metric Graphs.” Mathematics
of Computation 93 (349): 2439–72.
https://doi.org/10.1090/mcom/3929.
Bolin, David, and Alexandre B. Simas. 2023.
rSPDE: Rational Approximations of Fractional
Stochastic Partial Differential Equations.
https://CRAN.R-project.org/package=rSPDE.
Bolin, David, Alexandre B. Simas, and Jonas Wallin. 2023a.
MetricGraph: Random Fields on Metric Graphs.
https://CRAN.R-project.org/package=MetricGraph.
———. 2023b.
“Statistical Inference for Gaussian Whittle-Matérn
Fields on Metric Graphs.” arXiv Preprint
arXiv:2304.10372.
https://doi.org/10.48550/arXiv.2304.10372.
———. 2024.
“Gaussian Whittle-Matérn Fields on Metric
Graphs.” Bernoulli 30 (2): 1611–39.
https://doi.org/10.3150/23-BEJ1647.
———. 2025.
“Markov Properties of Gaussian Random Fields on Compact
Metric Graphs.” Bernoulli.
https://doi.org/10.48550/arXiv.2304.03190.
Bolin, David, Alexandre B. Simas, and Zhen Xiong. 2024.
“Covariance-Based Rational Approximations of Fractional SPDEs for
Computationally Efficient Bayesian Inference.” Journal of
Computational and Graphical Statistics 33 (1): 64–74.
https://doi.org/10.1080/10618600.2023.2231051.
De Coninck, Arne, Bernard De Baets, Drosos Kourounis, Fabio Verbosio,
Olaf Schenk, Steven Maenhout, and Jan Fostier. 2016.
“Needles: Toward Large-Scale Genomic Prediction with
Marker-by-Environment Interaction.” Genetics 203 (1):
543–55.
https://doi.org/10.1534/genetics.115.179887.
Fellows, Ian, and Jan-Peter Stotz. 2025.
OpenStreetMap:
Access to Open Street Map Raster Images.
https://doi.org/10.32614/CRAN.package.OpenStreetMap.
Garnier, Simon, Ross, Noam, Rudis, Robert, Camargo, et al. 2024.
viridis(Lite) - Colorblind-Friendly
Color Maps for r.
https://doi.org/10.5281/zenodo.4679423.
Hester, Jim, and Jennifer Bryan. 2024.
glue: Interpreted String Literals.
https://glue.tidyverse.org/.
J, Lemon. 2006. “Plotrix: A Package in the Red Light
District of r.” R-News 6 (4): 8–12.
Kahle, David, and Hadley Wickham. 2013.
“ggmap: Spatial Visualization with Ggplot2.”
The R Journal 5 (1): 144–61.
https://journal.r-project.org/archive/2013-1/kahle-wickham.pdf.
Kassambara, Alboukadel. 2026.
ggpubr:
“ggplot2” Based Publication
Ready Plots.
https://doi.org/10.32614/CRAN.package.ggpubr.
Kaye, Matt, Bob Rudis, Andrie de Vries, and Jonathan Sidi. 2025.
slackr: Send Messages, Images, r Objects
and Files to “Slack” Channels/Users.
https://github.com/mrkaye97/slackr.
Kourounis, D., A. Fuchs, and O. Schenk. 2018.
“Towards the Next
Generation of Multiperiod Optimal Power Flow Solvers.” IEEE
Transactions on Power Systems PP (99): 1–10.
https://doi.org/10.1109/TPWRS.2017.2789187.
Lindgren, Finn, and Håvard Rue. 2015.
“Bayesian Spatial Modelling
with R-INLA.” Journal of
Statistical Software 63 (19): 1–25.
http://www.jstatsoft.org/v63/i19/.
Lindgren, Finn, Håvard Rue, and Johan Lindström. 2011. “An
Explicit Link Between Gaussian Fields and
Gaussian Markov Random Fields: The Stochastic
Partial Differential Equation Approach (with Discussion).”
Journal of the Royal Statistical Society B 73 (4): 423–98.
Martins, Thiago G., Daniel Simpson, Finn Lindgren, and Håvard Rue. 2013.
“Bayesian Computing with INLA: New
Features.” Computational Statistics and Data Analysis
67: 68–83.
Meschiari, Stefano. 2026.
Latex2exp: Use LaTeX Expressions in
Plots.
https://doi.org/10.32614/CRAN.package.latex2exp.
Müller, Kirill. 2020.
here: A Simpler
Way to Find Your Files.
https://doi.org/10.32614/CRAN.package.here.
Pebesma, Edzer. 2018.
“Simple Features for R:
Standardized Support for Spatial Vector Data.”
The R Journal 10 (1): 439–46.
https://doi.org/10.32614/RJ-2018-009.
Pebesma, Edzer J., and Roger Bivand. 2005.
“Classes and Methods
for Spatial Data in R.” R News 5 (2): 9–13.
https://CRAN.R-project.org/doc/Rnews/.
Pebesma, Edzer, and Roger Bivand. 2023.
Spatial
Data Science: With applications in R.
Chapman and
Hall/CRC.
https://doi.org/10.1201/9780429459016.
Pedersen, Thomas Lin. 2025.
patchwork:
The Composer of Plots.
https://doi.org/10.32614/CRAN.package.patchwork.
R Core Team. 2025.
R: A Language and Environment for
Statistical Computing. Vienna, Austria: R Foundation for
Statistical Computing.
https://www.R-project.org/.
Rue, Håvard, Sara Martino, and Nicholas Chopin. 2009. “Approximate
Bayesian Inference for Latent Gaussian Models
Using Integrated Nested Laplace Approximations (with
Discussion).” Journal of the Royal Statistical Society B
71: 319–92.
Rue, Håvard, Andrea I. Riebler, Sigrunn H. Sørbye, Janine B. Illian,
Daniel P. Simpson, and Finn K. Lindgren. 2017.
“Bayesian Computing
with INLA: A Review.” Annual
Reviews of Statistics and Its Applications 4 (March): 395–421.
http://arxiv.org/abs/1604.00860.
Sharpsteen, Charlie, and Cameron Bracken. 2023.
tikzDevice: R Graphics Output in LaTeX
Format.
https://doi.org/10.32614/CRAN.package.tikzDevice.
Sievert, Carson. 2020.
Interactive Web-Based Data Visualization with
r, Plotly, and Shiny. Chapman; Hall/CRC.
https://plotly-r.com.
Ushey, Kevin, JJ Allaire, and Yuan Tang. 2025.
reticulate: Interface to
“Python”.
https://doi.org/10.32614/CRAN.package.reticulate.
Ushey, Kevin, and Hadley Wickham. 2026.
renv: Project Environments.
https://doi.org/10.32614/CRAN.package.renv.
Verbosio, Fabio, Arne De Coninck, Drosos Kourounis, and Olaf Schenk.
2017.
“Enhancing the Scalability of Selected Inversion
Factorization Algorithms in Genomic Prediction.” Journal of
Computational Science 22 (Supplement C): 99–108.
https://doi.org/10.1016/j.jocs.2017.08.013.
Wickham, Hadley. 2007.
“Reshaping Data with the reshape Package.” Journal of
Statistical Software 21 (12): 1–20.
http://www.jstatsoft.org/v21/i12/.
Wickham, Hadley, Mara Averick, Jennifer Bryan, Winston Chang, Lucy
D’Agostino McGowan, Romain François, Garrett Grolemund, et al. 2019.
“Welcome to the tidyverse.”
Journal of Open Source Software 4 (43): 1686.
https://doi.org/10.21105/joss.01686.
Wickham, Hadley, Thomas Lin Pedersen, and Dana Seidel. 2025.
scales: Scale Functions for Visualization.
https://scales.r-lib.org.
Wilke, Claus O. 2025.
cowplot:
Streamlined Plot Theme and Plot Annotations for “ggplot2”.
https://doi.org/10.32614/CRAN.package.cowplot.
Wilke, Claus O., and Brenton M. Wiernik. 2022.
ggtext: Improved Text Rendering Support for
“ggplot2”.
https://doi.org/10.32614/CRAN.package.ggtext.
Xie, Yihui. 2014. “knitr: A
Comprehensive Tool for Reproducible Research in R.”
In Implementing Reproducible Computational Research, edited by
Victoria Stodden, Friedrich Leisch, and Roger D. Peng. Chapman;
Hall/CRC.
———. 2015.
Dynamic Documents with R and Knitr. 2nd
ed. Boca Raton, Florida: Chapman; Hall/CRC.
https://yihui.org/knitr/.
———. 2025.
knitr: A General-Purpose
Package for Dynamic Report Generation in R.
https://yihui.org/knitr/.
Xie, Yihui, J. J. Allaire, and Garrett Grolemund. 2018.
R Markdown:
The Definitive Guide. Boca Raton, Florida: Chapman; Hall/CRC.
https://bookdown.org/yihui/rmarkdown.
Xie, Yihui, Christophe Dervieux, and Emily Riederer. 2020.
R
Markdown Cookbook. Boca Raton, Florida: Chapman; Hall/CRC.
https://bookdown.org/yihui/rmarkdown-cookbook.
Yuan, Yuan, Bachl, Fabian E., Lindgren, Finn, Borchers, et al. 2017.
“Point Process Models for Spatio-Temporal Distance Sampling Data
from a Large-Scale Survey of Blue Whales.” Ann. Appl.
Stat. 11 (4): 2270–97.
https://doi.org/10.1214/17-AOAS1078.
LS0tCnRpdGxlOiAiUGVNUyAxLCBjb3ZhcmlhdGVzIGFuZCBwcmVwcm9jZXNzaW5nIgpkYXRlOiAiTGFzdCBtb2RpZmllZDogYHIgZm9ybWF0KFN5cy50aW1lKCksICclZC0lbS0lWS4nKWAiCm91dHB1dDoKICBodG1sX2RvY3VtZW50OgogICAgbWF0aGpheDogImh0dHBzOi8vY2RuLmpzZGVsaXZyLm5ldC9ucG0vbWF0aGpheEAzL2VzNS90ZXgtbW1sLWNodG1sLmpzIgogICAgaGlnaGxpZ2h0OiBweWdtZW50cwogICAgdGhlbWU6IGZsYXRseQogICAgY29kZV9mb2xkaW5nOiBoaWRlICMgY2xhc3Muc291cmNlID0gImZvbGQtaGlkZSIgdG8gaGlkZSBjb2RlIGFuZCBhZGQgYSBidXR0b24gdG8gc2hvdyBpdAogICAgZGZfcHJpbnQ6IHBhZ2VkCiAgICB0b2M6IHRydWUKICAgIHRvY19mbG9hdDoKICAgICAgY29sbGFwc2VkOiB0cnVlCiAgICAgIHNtb290aF9zY3JvbGw6IHRydWUKICAgIG51bWJlcl9zZWN0aW9uczogdHJ1ZQogICAgZmlnX2NhcHRpb246IHRydWUKICAgIGNvZGVfZG93bmxvYWQ6IHRydWUKICAgIGNzczogdmlzdWFsLmNzcwphbHdheXNfYWxsb3dfaHRtbDogdHJ1ZQpiaWJsaW9ncmFwaHk6IAogIC0gcmVmZXJlbmNlcy5iaWIKICAtIGdyYXRlZnVsLXJlZnMuYmliCmhlYWRlci1pbmNsdWRlczoKICAtIFxuZXdjb21tYW5ke1xhcn17XG1hdGhiYntSfX0KICAtIFxuZXdjb21tYW5ke1xsbGF2fVsxXXtcbGVmdFx7IzFccmlnaHRcfX0KICAtIFxuZXdjb21tYW5ke1xwYXJlfVsxXXtcbGVmdCgjMVxyaWdodCl9CiAgLSBcbmV3Y29tbWFuZHtcTmNhbH17XG1hdGhjYWx7Tn19CiAgLSBcbmV3Y29tbWFuZHtcVmNhbH17XG1hdGhjYWx7Vn19CiAgLSBcbmV3Y29tbWFuZHtcRWNhbH17XG1hdGhjYWx7RX19CiAgLSBcbmV3Y29tbWFuZHtcV2NhbH17XG1hdGhjYWx7V319CiAgLSBcbmV3Y29tbWFuZHtcYWxtb3N0ZXZlcnl3aGVyZX17XG1hdGhybXthLmUufVw7fQotLS0KCkdvIGJhY2sgdG8gdGhlIFtDb250ZW50c10oYWJvdXQuaHRtbCkgcGFnZS4KCjxkaXYgc3R5bGU9ImNvbG9yOiAjMmMzZTUwOyB0ZXh0LWFsaWduOiByaWdodDsiPgoqKioqKioqKiAgCjxzdHJvbmc+UHJlc3MgU2hvdyB0byByZXZlYWwgdGhlIGNvZGUgY2h1bmtzLjwvc3Ryb25nPiAgCgoqKioqKioqKgo8L2Rpdj4KCgoKR28gYmFjayB0byB0aGUgW0Fib3V0IHBhZ2VdKGFib3V0Lmh0bWwpLgoKCjo6OiB7LmN1c3RvbS1ib3h9ClRoaXMgdmlnbmV0dGUgcHJvY2Vzc2VzIGhhbGYgb2YgdGhlIFBlTVMgZGF0YXNldCwgc2hvd3Mgc29tZSBpbGx1c3RyYXRpb25zLCBhbmQgcHJvZHVjZXMgWyoqYHBlbXNfcmVwbDFfZGF0YS5SRGF0YWAqKl0oaHR0cHM6Ly9naXRodWIuY29tL2xlbmlucmFmYWVscmllcmFzZWd1cmEvR1dNRi9ibG9iL21haW4vZGF0YV9maWxlcy9wZW1zX3JlcGwxX2RhdGEuUkRhdGEpLCB3aGljaCBpcyBhIGZpbGUgdGhhdCBjb250YWlucyB0aGUgZ3JhcGggd2l0aCBkYXRhIGFuZCBhIGNvdmFyaWF0ZSBkZWZpbmVkIG9uIHRoZSBtZXNoLiBUbyBzZWUgaG93IHRoZSBkYXRhIGlzIG1vZGVsZWQsIGdvIHRvIFtwZW1zX3JlcGwyLmh0bWxdKHBlbXNfcmVwbDIuaHRtbCkuCjo6OgoKCkxldCB1cyBzZXQgc29tZSBnbG9iYWwgb3B0aW9ucyBmb3IgYWxsIGNvZGUgY2h1bmtzIGluIHRoaXMgZG9jdW1lbnQuCgoKYGBge3J9CiMgQ3JlYXRlIGEgY2xpcGJvYXJkIGJ1dHRvbiBvbiB0aGUgcmVuZGVyZWQgSFRNTCBwYWdlCnNvdXJjZShoZXJlOjpoZXJlKCJjbGlwYm9hcmQuUiIpKTsgY2xpcGJvYXJkCiMgU2V0IHNlZWQgZm9yIHJlcHJvZHVjaWJpbGl0eQpzZXQuc2VlZCgxOTgyKSAKIyBTZXQgZ2xvYmFsIG9wdGlvbnMgZm9yIGFsbCBjb2RlIGNodW5rcwprbml0cjo6b3B0c19jaHVuayRzZXQoCiAgIyBEaXNhYmxlIG1lc3NhZ2VzIHByaW50ZWQgYnkgUiBjb2RlIGNodW5rcwogIG1lc3NhZ2UgPSBGQUxTRSwgICAgCiAgIyBEaXNhYmxlIHdhcm5pbmdzIHByaW50ZWQgYnkgUiBjb2RlIGNodW5rcwogIHdhcm5pbmcgPSBGQUxTRSwgICAgCiAgIyBTaG93IFIgY29kZSB3aXRoaW4gY29kZSBjaHVua3MgaW4gb3V0cHV0CiAgZWNobyA9IFRSVUUsICAgICAgICAKICAjIEluY2x1ZGUgYm90aCBSIGNvZGUgYW5kIGl0cyByZXN1bHRzIGluIG91dHB1dAogIGluY2x1ZGUgPSBUUlVFLCAgICAgCiAgIyBFdmFsdWF0ZSBSIGNvZGUgY2h1bmtzCiAgZXZhbCA9IEZBTFNFLCAgICAgICAKICAjIEVuYWJsZSBjYWNoaW5nIG9mIFIgY29kZSBjaHVua3MgZm9yIGZhc3RlciByZW5kZXJpbmcKICBjYWNoZSA9IEZBTFNFLCAgICAgIAogICMgQWxpZ24gZmlndXJlcyBpbiB0aGUgY2VudGVyIG9mIHRoZSBvdXRwdXQKICBmaWcuYWxpZ24gPSAiY2VudGVyIiwKICAjIEVuYWJsZSByZXRpbmEgZGlzcGxheSBmb3IgaGlnaC1yZXNvbHV0aW9uIGZpZ3VyZXMKICByZXRpbmEgPSAyLAogICMgU2hvdyBlcnJvcnMgaW4gdGhlIG91dHB1dCBpbnN0ZWFkIG9mIHN0b3BwaW5nIHJlbmRlcmluZwogIGVycm9yID0gVFJVRSwKICAjIERvIG5vdCBjb2xsYXBzZSBjb2RlIGFuZCBvdXRwdXQgaW50byBhIHNpbmdsZSBibG9jawogIGNvbGxhcHNlID0gRkFMU0UKKQojIFN0YXJ0IHRoZSBmaWd1cmUgY291bnRlcgpmaWdfY291bnQgPC0gMAojIERlZmluZSB0aGUgY2FwdGlvbmVyIGZ1bmN0aW9uCmNhcHRpb25lciA8LSBmdW5jdGlvbihjYXB0aW9uKSB7CiAgZmlnX2NvdW50IDw8LSBmaWdfY291bnQgKyAxCiAgcGFzdGUwKCJGaWd1cmUgIiwgZmlnX2NvdW50LCAiOiAiLCBjYXB0aW9uKQp9CiMgRGVmaW5lIHRoZSBmdW5jdGlvbiB0byB0cnVuY2F0ZSBhIG51bWJlciB0byB0d28gZGVjaW1hbCBwbGFjZXMKdHJ1bmNhdGVfdG9fdHdvIDwtIGZ1bmN0aW9uKHgpIHsKICB0cnVuY2F0ZWQgPC0gZmxvb3IoeCAqIDEwMCkgLyAxMDAKICBzcHJpbnRmKCIlLjJmIiwgdHJ1bmNhdGVkKQp9CmBgYAoKCkJlbG93IHdlIGxvYWQgdGhlIG5lY2Vzc2FyeSBsaWJyYXJpZXMuCgpgYGB7ciwgZXZhbCA9IFRSVUV9CmxpYnJhcnkoSU5MQSkKbGlicmFyeShpbmxhYnJ1KQpsaWJyYXJ5KHJTUERFKQpsaWJyYXJ5KE1ldHJpY0dyYXBoKQoKbGlicmFyeShkcGx5cikKbGlicmFyeShwbG90bHkpCmxpYnJhcnkoc2NhbGVzKQpsaWJyYXJ5KHBhdGNod29yaykKCmxpYnJhcnkoZ2dwbG90MikKbGlicmFyeShjb3dwbG90KQpsaWJyYXJ5KGdncHVicikgI2Fubm90YXRlX2ZpZ3VyZSgpCmxpYnJhcnkoZ3JpZCkgI3RleHRHcm9iKCkKbGlicmFyeShnZ21hcCkKCmxpYnJhcnkodmlyaWRpcykKbGlicmFyeShPcGVuU3RyZWV0TWFwKQoKCmxpYnJhcnkodGlkeXIpCmxpYnJhcnkoc2YpCgpsaWJyYXJ5KGhlcmUpCmxpYnJhcnkocm1hcmtkb3duKQpsaWJyYXJ5KGdyYXRlZnVsKSAjIENpdGUgYWxsIGxvYWRlZCBwYWNrYWdlcwoKbGlicmFyeShzbGFja3IpCnNvdXJjZSgia2V5cy5SIikKc2xhY2tyX3NldHVwKHRva2VuID0gdG9rZW4pICMgdG9rZW4gY29tZXMgZnJvbSBrZXlzLlIKYGBgCgoKYGBge3J9CmNhcHR1cmUub3V0cHV0KAogIGtuaXRyOjpwdXJsKGhlcmU6OmhlcmUoImZ1bmN0aW9uYWxpdHkxLlJtZCIpLCBvdXRwdXQgPSBoZXJlOjpoZXJlKCJmdW5jdGlvbmFsaXR5MS5SIikpLAogIGZpbGUgPSBoZXJlOjpoZXJlKCJvbGQvcHVybF9sb2cudHh0IikKKQpzb3VyY2UoaGVyZTo6aGVyZSgiZnVuY3Rpb25hbGl0eTEuUiIpKQpgYGAKCgpCZWxvdyB3ZSBkZWZpbmUgZnVuY3Rpb24gYGdldHNfc3VtbWFyeV9wYXJhbWV0ZXJzKClgIHRvIGV4dHJhY3QgdGhlIHN1bW1hcnkgb2YgdGhlIHBhcmFtZXRlcnMgb2YgdGhlIG1vZGVsLgoKCgpgYGB7ciwgZXZhbCA9IFRSVUV9CmdldHNfc3VtbWFyeV9wYXJhbWV0ZXJzIDwtIGZ1bmN0aW9uKGZpdCwgbW9kZWwpIHsKICBwYXJhbV9zcGRlIDwtIHN1bW1hcnkocnNwZGUucmVzdWx0KGZpdCwgImZpZWxkIiwgbW9kZWwsIHBhcmFtZXRlcml6YXRpb24gPSAic3BkZSIpKQogIHBhcmFtX21hdGVybiA8LSBzdW1tYXJ5KHJzcGRlLnJlc3VsdChmaXQsICJmaWVsZCIsIG1vZGVsLCBwYXJhbWV0ZXJpemF0aW9uID0gIm1hdGVybiIpKQogIHBhcmFtX2ZpeGVkIDwtIGZpdCRzdW1tYXJ5LmZpeGVkWywxOjZdCiAgbWFyZ2luYWwucG9zdGVyaW9yLnNpZ21hX2UgPSBpbmxhLnRtYXJnaW5hbCgKICAgIGZ1biA9IGZ1bmN0aW9uKHgpIGV4cCgteC8yKSwgCiAgICBtYXJnaW5hbCA9IGZpdFtbImludGVybmFsLm1hcmdpbmFscy5oeXBlcnBhciJdXVtbIkxvZyBwcmVjaXNpb24gZm9yIHRoZSBHYXVzc2lhbiBvYnNlcnZhdGlvbnMiXV0pCiAgcXVhbnQuc2lnbWFfZSA8LSBjYXB0dXJlLm91dHB1dCh7cmVzdWx0X3RtcCA8LSBpbmxhLnptYXJnaW5hbChtYXJnaW5hbC5wb3N0ZXJpb3Iuc2lnbWFfZSl9LCBmaWxlID0gIi9kZXYvbnVsbCIpIAogIHF1YW50LnNpZ21hX2UgPC0gcmVzdWx0X3RtcAogIHN0YXRpc3RpY3Muc2lnbWFfZSA8LSB1bmxpc3QocXVhbnQuc2lnbWFfZSlbYygxLDIsMyw1LDcpXQogIG1vZGUuc2lnbWFfZSA8LSBpbmxhLm1tYXJnaW5hbChtYXJnaW5hbC5wb3N0ZXJpb3Iuc2lnbWFfZSkKICBhbGxwYXJhbXMgPC0gcmJpbmQocGFyYW1fZml4ZWQsIHBhcmFtX3NwZGUsIHBhcmFtX21hdGVybiwgYyhzdGF0aXN0aWNzLnNpZ21hX2UsIG1vZGUuc2lnbWFfZSkpCiAgcm93bmFtZXMoYWxscGFyYW1zKVtucm93KGFsbHBhcmFtcyldIDwtICJzaWdtYV9lIgogIHJldHVybihhbGxwYXJhbXMpCn0KYGBgCgoKCjwhLS0gT2JqZWN0IGBwZW1zX3JlcGxgIGJlbG93IGlzIGF2YWlsYWJsZSBhZnRlciBsb2FkaW5nIGBNZXRyaWNHcmFwaGAgbGlicmFyeS4gLS0+Cgo8IS0tIGBgYHtyfSAtLT4KPCEtLSAjIExvYWQgdGhlIGRhdGEgLS0+CjwhLS0gcGVtc19kYXRhIDwtIHBlbXNfcmVwbCRkYXRhIC0tPgo8IS0tICMgU2VsZWN0IHRoZSBsb2NhdGlvbnMsIHdoaWNoIGFyZSBjb21tb24gZm9yIGFsbCByZXBsaWNhdGVzIC0tPgo8IS0tIFB0RV9hbGwgPC0gcGVtc19kYXRhWzE6MzE5LCBjKCJlZGdlX251bWJlciIsICJkaXN0YW5jZV9vbl9lZGdlIildIC0tPgo8IS0tICMgU2VsZWN0IHRoZSByZXNwb25zZSB2YXJpYWJsZSBhbmQgdHJhbnNmb3JtIGl0IHRvIGEgbWF0cml4IC0tPgo8IS0tIFlfYWxsIDwtIG1hdHJpeChwZW1zX2RhdGEkeSwgbnJvdyA9IDI2LCBuY29sID0gMzE5LCBieXJvdyA9IFRSVUUpIC0tPgo8IS0tICMgUmVtb3ZlIGxvY2F0aW9ucywgdGhvc2Ugd2l0aCB0aGUgc2FtZSB2YWx1ZSBpbiB0aGUgZmlyc3QgMTMgcmVwbGljYXRlcyAtLT4KPCEtLSBhbGxfc2FtZSA8LSBmdW5jdGlvbihjb2wpIHtsZW5ndGgodW5pcXVlKGNvbFsxOjEzXSkpID09IDF9IC0tPgo8IS0tIGNvbHNfdG9fa2VlcCA8LSBhcHBseShZX2FsbCwgMiwgZnVuY3Rpb24oY29sKSAhYWxsX3NhbWUoY29sKSkgLS0+CjwhLS0gWV9sb2MgPC0gWV9hbGxbLCBjb2xzX3RvX2tlZXBdIC0tPgo8IS0tIFB0RSA8LSBQdEVfYWxsW2NvbHNfdG9fa2VlcCxdIC0tPgo8IS0tICMgQ2hlY2sgdGhlIGRpbWVuc2lvbnMgLS0+CjwhLS0gUHRFIHw+IGRpbSgpIC0tPgo8IS0tIFlfbG9jIHw+IGRpbSgpIC0tPgo8IS0tIGBgYCAtLT4KCjwhLS0gSGFsZiBvZiB0aGUgcmVwbGljYXRlcyBhcmUgdXNlZCB0byBjb21wdXRlIHN1bW1hcnkgc3RhdGlzdGljcyAoYFlfbG9nc3RkYCBhbmQgYFlfbWVhbmApIGFuZCB0aGUgb3RoZXIgaGFsZiAoYFlgKSBhcmUgdXNlZCB0byBmaXQgdGhlIG1vZGVsLiAtLT4KCjwhLS0gYGBge3J9IC0tPgo8IS0tICMgRmlyc3QgaGFsZiBvZiB0aGUgcmVwbGljYXRlcyBmb3Igc3VtbWFyeSBzdGF0aXN0aWNzIC0tPgo8IS0tIFlfZm9yX3N1bW1hcnkgPC0gWV9sb2NbMToxMyxdIC0tPgo8IS0tICMgUmVzZXJ2ZSB0aGUgc2Vjb25kIGhhbGYgZm9yIGZpdHRpbmcgcmVwbGljYXRlIG1vZGVsIC0tPgo8IS0tIFkgPC0gWV9sb2NbMTQ6MjYsXSAtLT4KPCEtLSAjIENvbXB1dGUgdGhlIGxvZyBzdGFuZGFyZCBkZXZpYXRpb24gYW5kIG1lYW4gLS0+CjwhLS0gWV9sb2dzdGQgPC0gYXBwbHkoWV9mb3Jfc3VtbWFyeSwgMiwgc2QpIHw+IGFzLnZlY3RvcigpIHw+IGxvZygpIC0tPgo8IS0tIFlfbWVhbiA8LSBhcHBseShZX2Zvcl9zdW1tYXJ5LCAyLCBtZWFuKSB8PiBhcy52ZWN0b3IoKSAtLT4KPCEtLSAjIENoZWNrIHRoZSBkaW1lbnNpb25zIC0tPgo8IS0tIFkgfD4gZGltKCkgLS0+CjwhLS0gYGBgIC0tPgoKCioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqCgpGb2xkZXIgYGRhdGEucGVtc2Agd2FzIGRvd25sb2FkZWQgZnJvbSB0aGlzIFtHaXRIdWIgcmVwb3NpdG9yeV0oaHR0cHM6Ly9naXRodWIuY29tL2RhdmlkYm9saW4vTWV0cmljR3JhcGgvdHJlZS83ZjRjOTAxNzFjNDBiOWJmN2VhMjllOWIwNDhjOTNkODg0NzdjNWQxL2V4YW1wbGVzL2RhdGEucGVtcykuCgpgYGB7cn0KTGluZXMgPC0gcmVhZF9zZihoZXJlKCJkYXRhX2ZpbGVzL2RhdGEucGVtcy9saW5lcy5zaHAiKSkKbGluZXMgPC0gYXNfU3BhdGlhbChMaW5lcykKCkV0ViA8LSByZWFkLmNzdihoZXJlKCJkYXRhX2ZpbGVzL2RhdGEucGVtcy9FLmNzdiIpLCBoZWFkZXIgPSBULCByb3cubmFtZXMgPSBOVUxMKQpQdEUgPC0gcmVhZC5jc3YoaGVyZSgiZGF0YV9maWxlcy9kYXRhLnBlbXMvUHRFLmNzdiIpLCBoZWFkZXIgPSBULCByb3cubmFtZXMgPSBOVUxMKQpQdEVbLDFdIDwtIFB0RVssMV0gKyAxClkgPC0gcmVhZC5jc3YoaGVyZSgiZGF0YV9maWxlcy9kYXRhLnBlbXMvWS5jc3YiKSwgaGVhZGVyID0gVCwgcm93Lm5hbWVzID0gTlVMTCkKWSA8LSBhcy5tYXRyaXgoWVssLTFdKQplZGdlX2xlbmd0aF9tIDwtIEV0VlssNF0KUHRFWywyXSA9IFB0RVssMl0vZWRnZV9sZW5ndGhfbVtQdEVbLDFdXQpgYGAKCk1hdHJpeCBgWWAgaGFzIGRpbWVuc2lvbiAkMjZcdGltZXMzMjUkLiBFYWNoIHJvdyBvZiBgWWAgY29ycmVzcG9uZHMgdG8gYSByZXBsaWNhdGUgKCQyNiQgcmVwbGljYXRlcyBpbiB0b3RhbCkgYW5kIGVhY2ggY29sdW1uIGNvcnJlc3BvbmRzIHRvIGEgbG9jYXRpb24gKCQzMjUkIGxvY2F0aW9ucyBpbiB0b3RhbCkgb24gdGhlIG5ldHdvcmsuCgoKV2UgcmVtb3ZlIGNvbHVtbnMgaW4gYFlgLCB0aG9zZSB3aXRoIHRoZSBzYW1lIHZhbHVlIGluIHRoZSBmaXJzdCAxMyByZXBsaWNhdGVzLiBXZSBhbHNvIHJlbW92ZSB0aGUgY29ycmVzcG9uZGluZyByb3dzIGluIGBQdEVgLgoKYGBge3J9CmFsbF9zYW1lIDwtIGZ1bmN0aW9uKGNvbCkgewogIGxlbmd0aCh1bmlxdWUoY29sWzE6MTNdKSkgPT0gMQp9CmNvbHNfdG9fa2VlcCA8LSBhcHBseShZLCAyLCBmdW5jdGlvbihjb2wpICFhbGxfc2FtZShjb2wpKQoKWSA8LSBZWywgY29sc190b19rZWVwXQpQdEUgPC0gUHRFW2NvbHNfdG9fa2VlcCxdCmBgYAoKCkhhbGYgb2YgdGhlIHJlcGxpY2F0ZXMgKGBZX2Zvcl9zdW1tYXJ5YCkgYXJlIHVzZWQgdG8gY29tcHV0ZSBgWV9sb2dzdGRgIGFuZCBgWV9tZWFuYCwgYW5kIHRoZSBvdGhlciBoYWxmIChgWWAgYWdhaW4gKSBhcmUgdXNlZCB0byBmaXQgYSByZXBsaWNhdGUgbW9kZWwgaW4gW3BlbXNfcmVwbDIuaHRtbF0ocGVtc19yZXBsMi5odG1sKS4KCmBgYHtyfQpzYW1wbGVkX251bWJlcnMgPC0gMToxMwpub3Rfc2FtcGxlZF9udW1iZXJzIDwtIDE0OjI2CllfZm9yX3N1bW1hcnkgPC0gWVtzYW1wbGVkX251bWJlcnMsXQpZX2xvZ3N0ZCA8LSBhcHBseShZX2Zvcl9zdW1tYXJ5LCAyLCBzZCkgfD4gYXMudmVjdG9yKCkgfD4gbG9nKCkKWV9tZWFuIDwtIGFwcGx5KFlfZm9yX3N1bW1hcnksIDIsIG1lYW4pIHw+IGFzLnZlY3RvcigpClkgPC0gWVtub3Rfc2FtcGxlZF9udW1iZXJzLF0Kc2F2ZShZX21lYW4sIGZpbGUgPSBoZXJlOjpoZXJlKCJkYXRhX2ZpbGVzL1lfbWVhbi5SRGF0YSIpKQpgYGAKCgoqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKioqKgoKCk9ic2VydmUgdGhhdCBgWV9sb2dzdGRgIGNvcnJlc3BvbmRzIHRvICRcbG9nJCBvZiB0aGUgc3RhbmRhcmQgZGV2aWF0aW9uIGF0IGVhY2ggbG9jYXRpb24uCgpCZWxvdyB3ZSBidWlsZCB0aGUgZ3JhcGggdXNpbmcgdGhlIGxpc3QgZWxlbWVudCBgZWRnZXNgIG9iamVjdCBmcm9tIGBwZW1zX3JlcGxgLgoKYGBge3J9CiMgQnVpbGQgdGhlIGdyYXBoCmdyYXBoIDwtIG1ldHJpY19ncmFwaCRuZXcoZWRnZXMgPSBwZW1zX3JlcGwkZWRnZXMsIGxvbmdsYXQgPSBUUlVFKQojIEFkZCB0aGUgb2JzZXJ2YXRpb25zCmdyYXBoJGFkZF9vYnNlcnZhdGlvbnMoZGF0YSA9IGRhdGEuZnJhbWUoeSA9IFlfbG9nc3RkLCAKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZGdlX251bWJlciA9IFB0RVssMV0sIAogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGRpc3RhbmNlX29uX2VkZ2UgPSBQdEVbLDJdKSwKICAgICAgICAgICAgICAgICAgICAgICBlZGdlX251bWJlciA9ICJlZGdlX251bWJlciIsCiAgICAgICAgICAgICAgICAgICAgICAgZGlzdGFuY2Vfb25fZWRnZSA9ICJkaXN0YW5jZV9vbl9lZGdlIiwKICAgICAgICAgICAgICAgICAgICAgICBkYXRhX2Nvb3JkcyA9ICJQdEUiLAogICAgICAgICAgICAgICAgICAgICAgIG5vcm1hbGl6ZWQgPSBUUlVFLCAKICAgICAgICAgICAgICAgICAgICAgICBjbGVhcl9vYnMgPSBUUlVFKQojIEJ1aWxkIHRoZSBtZXNoCmdyYXBoJGJ1aWxkX21lc2goaCA9IDAuMDUpIApgYGAKCkJlbG93IHdlIGZpdCAkXGxvZyhcbWF0aHJte3NkfShzX2kpKVxzaW0gTihcYmV0YV8wK3Uoc19pKSwgXHNpZ21hX2VeMikkLgoKCgpgYGB7ciwgZXZhbCA9IEZBTFNFfQojIEJ1aWxkIHRoZSBtb2RlbApyc3BkZV9tb2RlbF9zdGF0X2xvZ3N0ZCA8LSByc3BkZS5tZXRyaWNfZ3JhcGgoZ3JhcGgsCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHBhcmFtZXRlcml6YXRpb24gPSAic3BkZSIpCiMgUHJlcGFyZSB0aGUgZGF0YSBmb3IgZml0dGluZwpkYXRhX3JzcGRlX2JydV9zdGF0IDwtIGdyYXBoX2RhdGFfcnNwZGUocnNwZGVfbW9kZWxfc3RhdF9sb2dzdGQsIGJydSA9IFRSVUUpCiMgRGVmaW5lIHRoZSBjb21wb25lbnQKY21wX3N0YXQgPC0geSB+IC0xICsKICBJbnRlcmNlcHQoMSkgKwogIGZpZWxkKGNiaW5kKC5lZGdlX251bWJlciwgLmRpc3RhbmNlX29uX2VkZ2UpLCBtb2RlbCA9IHJzcGRlX21vZGVsX3N0YXRfbG9nc3RkKQojIEZpdCB0aGUgbW9kZWwKcnNwZGVfZml0X3N0YXRfbG9nc3RkIDwtCiAgYnJ1KGNtcF9zdGF0LAogICAgICBkYXRhID0gZGF0YV9yc3BkZV9icnVfc3RhdFtbImRhdGEiXV0sCiAgICAgIGZhbWlseSA9ICJnYXVzc2lhbiIsCiAgICAgIG9wdGlvbnMgPSBsaXN0KHZlcmJvc2UgPSBGQUxTRSkKICApCnNhdmUocnNwZGVfZml0X3N0YXRfbG9nc3RkLCByc3BkZV9tb2RlbF9zdGF0X2xvZ3N0ZCwgZmlsZSA9IGhlcmU6OmhlcmUoImRhdGFfZmlsZXMvcnNwZGVfZml0X3N0YXRfYW5kX21vZGVsX2xvZ3N0ZC5SRGF0YSIpKQpgYGAKCgpgYGB7ciwgZXZhbCA9IFRSVUV9CmxvYWQoaGVyZTo6aGVyZSgiZGF0YV9maWxlcy9yc3BkZV9maXRfc3RhdF9hbmRfbW9kZWxfbG9nc3RkLlJEYXRhIikpCiMgU3VtbWFyaXplIHRoZSByZXN1bHRzCnN1bW1hcnkocnNwZGVfZml0X3N0YXRfbG9nc3RkKQpnZXRzX3N1bW1hcnlfcGFyYW1ldGVycyhyc3BkZV9maXRfc3RhdF9sb2dzdGQsIHJzcGRlX21vZGVsX3N0YXRfbG9nc3RkKQpgYGAKCgpXZSBub3cgY29tcHV0ZSB0aGUga3JpZ2luZyBwcmVkaWN0b3IgJEUodShzX2kpfFxsb2coXG1hdGhybXtzZH0oc19pKSkpJCAoZGVub3RlZCBhcyBgY292YCBiZWxvdykgYW5kIHN0YW5kYXJkaXplIGl0LgoKYGBge3J9CiMgUHJlZGljdGlvbiBsb2NhdGlvbnMKZGF0YV9wcmRfbGlzdF9tZXNoIDwtIGdyYXBoJGdldF9tZXNoX2xvY2F0aW9ucyhicnUgPSBUUlVFKQojIENvbXB1dGUgdGhlIGtyaWdpbmcgcHJlZGljdG9yCnlfcHJlZCA8LSBwcmVkaWN0KHJzcGRlX2ZpdF9zdGF0X2xvZ3N0ZCwgbmV3ZGF0YSA9IGRhdGFfcHJkX2xpc3RfbWVzaCwgfkludGVyY2VwdCArIGZpZWxkKQpjb3ZfYWxtb3N0IDwtIHlfcHJlZCRtZWFuCiMgU3RhbmRhcmRpemUgdGhlIGtyaWdpbmcgcHJlZGljdG9yIG9mIHRoZSBsb2cgc3RhbmRhcmQgZGV2aWF0aW9uCmNvdiA8LSAoY292X2FsbW9zdCAtIG1lYW4oY292X2FsbW9zdCkpL3NkKGNvdl9hbG1vc3QpCmBgYAoKQmVsb3cgd2UgcGxvdCB0aGUgcmVzdWx0cy4gV2UgZmlyc3QgZ2VuZXJhdGUgdHdvIG1hcHMsIG9uZSBmb3Igem9vbSBsZXZlbCAxMiBhbmQgdGhlIG90aGVyIGZvciB6b29tIGxldmVsIDEzLiAKCgoKCmBgYHtyfQojIEV4dHJhY3QgdGhlIEV1Y2xpZGVhbiBjb29yZGluYXRlcyBvZiB0aGUgbWVzaCBwb2ludHMKeHlwb2ludHMgPC0gZ3JhcGgkbWVzaCRWCiMgRXh0cmFjdCB0aGUgcmFuZ2Ugb2YgdGhlIGNvb3JkaW5hdGVzIAp4X2xlZnQgPC0gcmFuZ2UoeHlwb2ludHNbLDFdKVsxXQp4X3JpZ2h0IDwtIHJhbmdlKHh5cG9pbnRzWywxXSlbMl0KeV9ib3R0b20gPC0gcmFuZ2UoeHlwb2ludHNbLDJdKVsxXQp5X3RvcCA8LSByYW5nZSh4eXBvaW50c1ssMl0pWzJdCmBgYAoKCmBgYHtyLCBldmFsID0gRkFMU0V9CiMgU2V0IHRoZSBzdHlsZSBvZiB0aGUgbWFwCnN0eWxlX3ZlY3RvciA8LSBjKGMoZmVhdHVyZSA9ICJyb2FkIiwgZWxlbWVudCA9ICJnZW9tZXRyeSIsIHZpc2liaWxpdHkgPSAib24iKSwKICAgICAgICAgICAgICAgICAgYygiJnN0eWxlPSIsIGZlYXR1cmUgPSAicG9pIiwgZWxlbWVudCA9ICJsYWJlbHMiLCB2aXNpYmlsaXR5ID0gIm9mZiIpLAogICAgICAgICAgICAgICAgICBjKCImc3R5bGU9IiwgZmVhdHVyZSA9ICJyb2FkIiwgZWxlbWVudCA9ICJsYWJlbHMiLCB2aXNpYmlsaXR5ID0gIm9mZiIpLAogICAgICAgICAgICAgICAgICBjKCImc3R5bGU9IiwgZmVhdHVyZSA9ICJhZG1pbmlzdHJhdGl2ZSIsIGVsZW1lbnQgPSAibGFiZWxzIiwgdmlzaWJpbGl0eSA9ICJvZmYiKSwgI25hbWUgb2YgcGxhY2VzCiAgICAgICAgICAgICAgICAgIGMoIiZzdHlsZT0iLCBmZWF0dXJlID0gInRyYW5zaXQiLCBlbGVtZW50ID0gImxhYmVscyIsIHZpc2liaWxpdHkgPSAib2ZmIiksCiAgICAgICAgICAgICAgICAgIGMoIiZzdHlsZT0iLCBmZWF0dXJlID0gImxhbmRzY2FwZSIsIGVsZW1lbnQgPSAiZ2VvbWV0cnkiLCB2aXNpYmlsaXR5ID0gIm9uIiksCiAgICAgICAgICAgICAgICAgIGMoIiZzdHlsZT0iLCBmZWF0dXJlID0gIndhdGVyIiwgZWxlbWVudCA9ICJnZW9tZXRyeSIsIHZpc2liaWxpdHkgPSAib24iKSkgCnJlZ2lzdGVyX3N0YWRpYW1hcHMoU3lzLmdldGVudigiR0dNQVBfU1RBRElBTUFQU19BUElfS0VZIikpCiMgT2J0YWluIHRoZSBtYXBzCnAxMiA8LSBnZ21hcChnZXRfc3RhZGlhbWFwKGJib3ggPSBjKGxlZnQgPSB4X2xlZnQsIGJvdHRvbSA9IHlfYm90dG9tLCByaWdodCA9IHhfcmlnaHQsIHRvcCA9IHlfdG9wKSwKICAgICAgICAgICAgICB6b29tID0gMTIsCiAgICAgICAgICAgICAgbWFwdHlwZSA9ICJzdGFtZW5fdG9uZXJfbGl0ZSIsIAogICAgICAgICAgICAgIHNjYWxlID0yLAogICAgICAgICAgICAgIHN0eWxlID0gc3R5bGVfdmVjdG9yLAogICAgICAgICAgICAgIGNvbG9yID0gImNvbG9yIikpICsgeGxhYihOVUxMKSArIHlsYWIoTlVMTCkKcDEzIDwtIGdnbWFwKGdldF9zdGFkaWFtYXAoYmJveCA9IGMobGVmdCA9IHhfbGVmdCwgYm90dG9tID0geV9ib3R0b20sIHJpZ2h0ID0geF9yaWdodCwgdG9wID0geV90b3ApLAogICAgICAgICAgICAgIHpvb20gPSAxMywKICAgICAgICAgICAgICBtYXB0eXBlID0gInN0YW1lbl90b25lcl9saXRlIiwgCiAgICAgICAgICAgICAgc2NhbGUgPTIsCiAgICAgICAgICAgICAgc3R5bGUgPSBzdHlsZV92ZWN0b3IsCiAgICAgICAgICAgICAgY29sb3IgPSAiY29sb3IiKSkgKyB4bGFiKE5VTEwpICsgeWxhYihOVUxMKQoKc2F2ZShwMTIsIHAxMywgZmlsZSA9IGhlcmU6OmhlcmUoImRhdGFfZmlsZXMvbWFwc196b29tMTJhbmQxM2Zyb21fc3RhZGlhLlJEYXRhIikpCmBgYAoKCmBgYHtyLCBldmFsID0gVFJVRX0KbG9hZChoZXJlOjpoZXJlKCJkYXRhX2ZpbGVzL21hcHNfem9vbTEyYW5kMTNmcm9tX3N0YWRpYS5SRGF0YSIpKQpgYGAKCgoKQmVsb3cgd2UgcGxvdCB0aGUgZmllbGQgb24gdG9wIG9mIHRoZSBtYXAuIFdlIGFsc28gYWRkIHRoZSBwb2ludCB2YWx1ZXMgb2YgdGhlIHN0YW5kYXJkIGRldmlhdGlvbiBvZiB0aGUgbG9nIHN0YW5kYXJkIGRldmlhdGlvbiB0byB0aGUgZ3JhcGggc28gd2UgY2FuIHBsb3QgaXQgb24gdG9wIG9mIHRoZSBtYXAgYW5kIHRoZSBmaWVsZC4KCmBgYHtyfQojIERlZmluZSB0aGUgY29sb3JzIGZvciB0aGUgd2luZG93cwpsb3dlcl9jb2xvciA8LSAiZGFya3JlZCIgICAKdXBwZXJfY29sb3IgPC0gImRhcmtibHVlIiAgCiMgRGVmaW5lIGNvb3JkaW5hdGVzIGZvciBzbWFsbCB3aW5kb3dzCmNvb3JkeF9sd3IxIDwtIC0xMjEuODc4CmNvb3JkeF91cHIxIDwtIC0xMjEuODI4CmNvb3JkeV9sd3IxIDwtIDM3LjMxNQpjb29yZHlfdXByMSA8LSAzNy4zNjUKCmNvb3JkeF9sd3IyPC0gLTEyMi4wNzUKY29vcmR4X3VwcjIgPC0gLTEyMi4wMjUKY29vcmR5X2x3cjIgPC0gMzcuMzY1CmNvb3JkeV91cHIyIDwtIDM3LjQxNQojIFBsb3QgdGhlIGZpZWxkIG9uIHRvcCBvZiB0aGUgbWFwCmYxMiA8LSBncmFwaCRwbG90X2Z1bmN0aW9uKFggPSBjb3YsIAogICAgICAgICAgICAgICAgICAgICAgICAgIHZlcnRleF9zaXplID0gMCwgCiAgICAgICAgICAgICAgICAgICAgICAgICAgcCA9IHAxMiwKICAgICAgICAgICAgICAgICAgICAgICAgICBlZGdlX3dpZHRoID0gMC41KSArIAogIHRoZW1lX21pbmltYWwoKSArIAogIHRoZW1lKHRleHQgPSBlbGVtZW50X3RleHQoZmFtaWx5ID0gIlBhbGF0aW5vIiksIAogICAgICAgIGF4aXMudGV4dCA9IGVsZW1lbnRfdGV4dChzaXplID0gOCksCiAgICAgICAgbGVnZW5kLnRleHQgPSBlbGVtZW50X3RleHQoc2l6ZSA9IDgpLAogICAgICAgIHBsb3QubWFyZ2luID0gdW5pdCgtMC40KmMoMSwwLDEsMSksICJjbSIpCiAgICAgICAgKSArCiAgbGFicyhjb2xvciA9ICIiLCB4ID0gIiIsIHkgPSAiIikgKwogIHhsaW0oeF9sZWZ0LCB4X3JpZ2h0KSArIAogIHlsaW0oeV9ib3R0b20sIHlfdG9wKQojIFBsb3QgdGhlIGZpZWxkIG9uIHRvcCBvZiB0aGUgbWFwCmYxMyA8LSBncmFwaCRwbG90X2Z1bmN0aW9uKFggPSBjb3YsIAogICAgICAgICAgICAgICAgICAgICAgICAgIHZlcnRleF9zaXplID0gMCwgCiAgICAgICAgICAgICAgICAgICAgICAgICAgcCA9IHAxMywKICAgICAgICAgICAgICAgICAgICAgICAgICBlZGdlX3dpZHRoID0gMC41KSArIAogIHRoZW1lX21pbmltYWwoKSArIAogIHRoZW1lKHRleHQgPSBlbGVtZW50X3RleHQoZmFtaWx5ID0gIlBhbGF0aW5vIiksIAogICAgICAgIGF4aXMudGV4dCA9IGVsZW1lbnRfdGV4dChzaXplID0gOCksCiAgICAgICAgbGVnZW5kLnRleHQgPSBlbGVtZW50X3RleHQoc2l6ZSA9IDgpLAogICAgICAgIHBsb3QubWFyZ2luID0gdW5pdCgtMC40KmMoMSwwLDEsMSksICJjbSIpCiAgICAgICAgKSArCiAgbGFicyhjb2xvciA9ICIiLCB4ID0gIiIsIHkgPSAiIikgKwogIHhsaW0oeF9sZWZ0LCB4X3JpZ2h0KSArIAogIHlsaW0oeV9ib3R0b20sIHlfdG9wKQoKIyBXZSBhZGQgdGhlIHBvaW50IHZhbHVlcyBvZiBzdGFuZGFyZCBkZXZpYXRpb24gb2YgdGhlIGxvZyBzdGFuZGFyZCBkZXZpYXRpb24gdG8gdGhlIGdyYXBoIHNvIHdlIGNhbiBwbG90IGl0CnN0YW5kYXJkaXplZF9ZX2xvZ3N0ZCA8LSAoWV9sb2dzdGQgLSBtZWFuKFlfbG9nc3RkKSkvc2QoWV9sb2dzdGQpCmdyYXBoJGFkZF9vYnNlcnZhdGlvbnMoZGF0YSA9IGRhdGEuZnJhbWUoeSA9IHN0YW5kYXJkaXplZF9ZX2xvZ3N0ZCwgCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWRnZV9udW1iZXIgPSBQdEVbLDFdLCAKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBkaXN0YW5jZV9vbl9lZGdlID0gUHRFWywyXSksIAogICAgICAgICAgICAgICAgICAgICAgIGVkZ2VfbnVtYmVyID0gImVkZ2VfbnVtYmVyIiwKICAgICAgICAgICAgICAgICAgICAgICBkaXN0YW5jZV9vbl9lZGdlID0gImRpc3RhbmNlX29uX2VkZ2UiLAogICAgICAgICAgICAgICAgICAgICAgIGRhdGFfY29vcmRzID0gIlB0RSIsCiAgICAgICAgICAgICAgICAgICAgICAgbm9ybWFsaXplZCA9IFRSVUUsIAogICAgICAgICAgICAgICAgICAgICAgIGNsZWFyX29icyA9IFRSVUUpCmcxMiA8LSBncmFwaCRwbG90KGRhdGEgPSAieSIsIHZlcnRleF9zaXplID0gMCwgcCA9IGYxMiwgZWRnZV93aWR0aCA9IDAsIGRhdGFfc2l6ZSA9IDEpICsgCiAgbGFicyhjb2xvciA9ICIiLCB4ID0gIiIsIHkgPSAiIikgKyAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMSwgeSA9IGNvb3JkeV9sd3IxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfbHdyMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIEJvdHRvbSBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjEsIHkgPSBjb29yZHlfdXByMSwgeGVuZCA9IGNvb3JkeF91cHIxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBUb3AgbGluZQogIGFubm90YXRlKCJzZWdtZW50IiwgeCA9IGNvb3JkeF9sd3IxLCB5ID0gY29vcmR5X2x3cjEsIHhlbmQgPSBjb29yZHhfbHdyMSwgeWVuZCA9IGNvb3JkeV91cHIxLCAKICAgICAgICAgICBsaW5ld2lkdGggPSAwLjQsIGNvbG9yID0gdXBwZXJfY29sb3IpICsgICMgTGVmdCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X3VwcjEsIHkgPSBjb29yZHlfbHdyMSwgeGVuZCA9IGNvb3JkeF91cHIxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBSaWdodCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X2x3cjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfdXByMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF9sd3IyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X3VwcjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikKZzEzIDwtIGdyYXBoJHBsb3QoZGF0YSA9ICJ5IiwgdmVydGV4X3NpemUgPSAwLCBwID0gZjEzLCBlZGdlX3dpZHRoID0gMCwgZGF0YV9zaXplID0gMSkgKyAKICBsYWJzKGNvbG9yID0gIiIsIHggPSAiIiwgeSA9ICIiKSArIAogIGFubm90YXRlKCJzZWdtZW50IiwgeCA9IGNvb3JkeF9sd3IxLCB5ID0gY29vcmR5X2x3cjEsIHhlbmQgPSBjb29yZHhfdXByMSwgeWVuZCA9IGNvb3JkeV9sd3IxLCAKICAgICAgICAgICBsaW5ld2lkdGggPSAwLjQsIGNvbG9yID0gdXBwZXJfY29sb3IpICsgICMgQm90dG9tIGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMSwgeSA9IGNvb3JkeV91cHIxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfdXByMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIFRvcCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjEsIHkgPSBjb29yZHlfbHdyMSwgeGVuZCA9IGNvb3JkeF9sd3IxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBMZWZ0IGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfdXByMSwgeSA9IGNvb3JkeV9sd3IxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfdXByMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIFJpZ2h0IGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfbHdyMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV91cHIyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X2x3cjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfdXByMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSAKCnIxIDwtIGcxMyArIHhsaW0oY29vcmR4X2x3cjEsIGNvb3JkeF91cHIxKSArIAogICAgICAgICAgICAgICAgICAgIHlsaW0oY29vcmR5X2x3cjEsIGNvb3JkeV91cHIxKSArIAogIHRoZW1lKGxlZ2VuZC5wb3NpdGlvbiA9ICJub25lIiwgCiAgICAgICAgcGxvdC5tYXJnaW4gPSB1bml0KC0wLjIqYygxLDEsMSwxKSwgImNtIikpCgpyMiA8LSBnMTMgKyB4bGltKGNvb3JkeF9sd3IyLCBjb29yZHhfdXByMikgKyAKICAgICAgICAgICAgICAgICAgICB5bGltKGNvb3JkeV9sd3IyLCBjb29yZHlfdXByMikgKyAKICB0aGVtZShsZWdlbmQucG9zaXRpb24gPSAibm9uZSIsIAogICAgICAgIHBsb3QubWFyZ2luID0gdW5pdCgtMC4yKmMoMSwxLDEsMSksICJjbSIpKQoKIyBBcnJhbmdlIHAyIGFuZCBwMyBob3Jpem9udGFsbHkKbGVmdF9jb2wgPC0gcGxvdF9ncmlkKHIyLCByMSwgbGFiZWxzID0gTlVMTCwgbmNvbCA9IDEsIG5yb3cgPSAyLCByZWxfaGVpZ2h0cyA9IGMoMSwxKSkKCiMgQ29tYmluZSB0aGUgdG9wIHJvdyB3aXRoIHAxIGluIGEgZ3JpZApjb21iaW5lZF9wbG90IDwtIHBsb3RfZ3JpZChsZWZ0X2NvbCwgZzEyLCBsYWJlbHMgPSBOVUxMLCBuY29sID0gMiwgcmVsX3dpZHRocyA9IGMoMSwyKSkgCmZpbmFsX3Bsb3QgPC0gYW5ub3RhdGVfZmlndXJlKGNvbWJpbmVkX3Bsb3QsIGxlZnQgPSB0ZXh0R3JvYigiTGF0aXR1ZGUiLCByb3QgPSA5MCwgdmp1c3QgPSAxLCBncCA9IGdwYXIoY2V4ID0gMC44KSksCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJvdHRvbSA9IHRleHRHcm9iKCJMb25naXR1ZGUiLCB2anVzdCA9IC0wLjUsIGdwID0gZ3BhcihjZXggPSAwLjgpKSkKZ2dzYXZlKGhlcmUoImRhdGFfZmlsZXMvc3RhbmRfbG9nX3NpZ21hXzMucG5nIiksIHdpZHRoID0gMTEuMiwgaGVpZ2h0ID0gNS40MywgcGxvdCA9IGZpbmFsX3Bsb3QsIGRwaSA9IDUwMCkKc2F2ZShmaW5hbF9wbG90LCBmaWxlID0gaGVyZSgiZGF0YV9maWxlcy9zdGFuZF9sb2dfc2lnbWFfMy5SRGF0YSIpKQpgYGAKCmBgYHtyLCBldmFsID0gVFJVRSwgZmlnLndpZHRoID0gMTEuMiwgZmlnLmhlaWdodCA9IDUuNDMsIGZpZy5jYXAgPSBjYXB0aW9uZXIoIlN0YW5kYXJkaXplZCBsb2dhcml0aG0gb2YgdGhlIHN0YW5kYXJkIGRldmlhdGlvbiBvZiB0aGUgZmlyc3QgcGFydCBvZiB0aGUgZGF0YS4gUG9pbnRzIHJlcHJlc2VudCB0aGUgcG9pbnQgdmFsdWUgYXQgZWFjaCBhdmFpbGFibGUgbG9jYXRpb24sIGFuZCBlZGdlcyByZXByZXNlbnQgdGhlICRcXG1hdGhybXtzdGQuY292fShzKSQgY292YXJpYXRlLiIpfQojIFByaW50IHRoZSBjb21iaW5lZCBwbG90CmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKGhlcmU6OmhlcmUoImRhdGFfZmlsZXMvc3RhbmRfbG9nX3NpZ21hXzMucG5nIikpCmBgYAoKCldlIHJlcGVhdCB0aGUgc2FtZSBwcm9jZXNzICBidXQgbm93IGZvciBgWV9tZWFuYC4gV2UgZmlyc3QgYWRkIHRoZSBvYnNlcnZhdGlvbnMgdG8gdGhlIGdyYXBoLgoKYGBge3J9CmdyYXBoJGFkZF9vYnNlcnZhdGlvbnMoZGF0YSA9IGRhdGEuZnJhbWUoeSA9IFlfbWVhbiwgCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWRnZV9udW1iZXIgPSBQdEVbLDFdLCAKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBkaXN0YW5jZV9vbl9lZGdlID0gUHRFWywyXSksCiAgICAgICAgICAgICAgICAgICAgICAgZWRnZV9udW1iZXIgPSAiZWRnZV9udW1iZXIiLAogICAgICAgICAgICAgICAgICAgICAgIGRpc3RhbmNlX29uX2VkZ2UgPSAiZGlzdGFuY2Vfb25fZWRnZSIsCiAgICAgICAgICAgICAgICAgICAgICAgZGF0YV9jb29yZHMgPSAiUHRFIiwKICAgICAgICAgICAgICAgICAgICAgICBub3JtYWxpemVkID0gVFJVRSwgCiAgICAgICAgICAgICAgICAgICAgICAgY2xlYXJfb2JzID0gVFJVRSkKYGBgCgpXZSBmaXQgdGhlIG1vZGVsICRcbWF0aHJte21lYW59KHNfaSlcc2ltIE4oXGJldGFfMCt1KHNfaSksIFxzaWdtYV9lXjIpJC4KCgoKYGBge3IsIGV2YWwgPSBGQUxTRX0KIyBCdWlsZCB0aGUgbW9kZWwKcnNwZGVfbW9kZWxfc3RhdF9tZWFuIDwtIHJzcGRlLm1ldHJpY19ncmFwaChncmFwaCwKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgcGFyYW1ldGVyaXphdGlvbiA9ICJzcGRlIikKIyBQcmVwYXJlIHRoZSBkYXRhIGZvciBmaXR0aW5nCmRhdGFfcnNwZGVfYnJ1X3N0YXQgPC0gZ3JhcGhfZGF0YV9yc3BkZShyc3BkZV9tb2RlbF9zdGF0X21lYW4sCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBicnUgPSBUUlVFKQojIERlZmluZSB0aGUgY29tcG9uZW50CmNtcF9zdGF0IDwtIHkgfiAtMSArCiAgSW50ZXJjZXB0KDEpICsKICBmaWVsZChjYmluZCguZWRnZV9udW1iZXIsIC5kaXN0YW5jZV9vbl9lZGdlKSwgbW9kZWwgPSByc3BkZV9tb2RlbF9zdGF0X21lYW4pCiMgRml0IHRoZSBtb2RlbApyc3BkZV9maXRfc3RhdF9tZWFuIDwtCiAgYnJ1KGNtcF9zdGF0LAogICAgICBkYXRhID0gZGF0YV9yc3BkZV9icnVfc3RhdFtbImRhdGEiXV0sCiAgICAgIGZhbWlseSA9ICJnYXVzc2lhbiIsCiAgICAgIG9wdGlvbnMgPSBsaXN0KHZlcmJvc2UgPSBGQUxTRSkKICApCnNhdmUocnNwZGVfZml0X3N0YXRfbWVhbiwgcnNwZGVfbW9kZWxfc3RhdF9tZWFuLCBmaWxlID0gaGVyZTo6aGVyZSgiZGF0YV9maWxlcy9yc3BkZV9maXRfc3RhdF9hbmRfbW9kZWxfbWVhbi5SRGF0YSIpKQpgYGAKCgpgYGB7ciwgZXZhbCA9IFRSVUV9CmxvYWQoaGVyZTo6aGVyZSgiZGF0YV9maWxlcy9yc3BkZV9maXRfc3RhdF9hbmRfbW9kZWxfbWVhbi5SRGF0YSIpKQojIFN1bW1hcml6ZSB0aGUgcmVzdWx0cwpzdW1tYXJ5KHJzcGRlX2ZpdF9zdGF0X21lYW4pCmdldHNfc3VtbWFyeV9wYXJhbWV0ZXJzKHJzcGRlX2ZpdF9zdGF0X21lYW4sIHJzcGRlX21vZGVsX3N0YXRfbWVhbikKYGBgCgoKQmVsb3cgd2UgY29tcHV0ZSB0aGUgcHJlZGljdGlvbnMgZm9yIHRoZSBtZXNoIGFuZCB0aGUgZGF0YSBsb2NhdGlvbi4KCmBgYHtyfQptZWFuX29uX21lc2hfcHJlZCA8LSBwcmVkaWN0KHJzcGRlX2ZpdF9zdGF0X21lYW4sIG5ld2RhdGEgPSBkYXRhX3ByZF9saXN0X21lc2gsIH5JbnRlcmNlcHQgKyBmaWVsZCkKY292X2Zvcl9tZWFuX3RvX3Bsb3QgPC0gbWVhbl9vbl9tZXNoX3ByZWQkbWVhbgoKbWVhbl9vbl9sb2NfcHJlZCA8LSBwcmVkaWN0KHJzcGRlX2ZpdF9zdGF0X21lYW4sIG5ld2RhdGEgPSByZW5hbWUoUHRFLCAuZWRnZV9udW1iZXIgPSBFLCAuZGlzdGFuY2Vfb25fZWRnZSA9IGxlbmd0aCksIH5JbnRlcmNlcHQgKyBmaWVsZCkKY292X2Zvcl9tZWFuIDwtIG1lYW5fb25fbG9jX3ByZWQkbWVhbgpzYXZlKGNvdl9mb3JfbWVhbl90b19wbG90LCBjb3ZfZm9yX21lYW4sIGZpbGUgPSBoZXJlOjpoZXJlKCJkYXRhX2ZpbGVzL2Nvdl9mb3JfbWVhbl90b19wbG90X2FuZF9jb3ZfZm9yX21lYW4uUkRhdGEiKSkKYGBgCgpCZWxvdyB3ZSBjaGVjayB0aGF0IHRoZSBwcmVkaWN0ZWQgbWVhbiBpcyBjbG9zZSB0byB0aGUgb2JzZXJ2ZWQgbWVhbi4KCmBgYHtyLCBldmFsID1UUlVFLCBmaWcud2lkdGggPSAyNCwgZmlnLmhlaWdodCA9IDQsIGZpZy5jYXAgPSBjYXB0aW9uZXIoIlByZWRpY3RlZCBtZWFuIGFuZCBvYnNlcnZlZCBtZWFuLiIpfQpsb2FkKGhlcmU6OmhlcmUoImRhdGFfZmlsZXMvWV9tZWFuLlJEYXRhIikpCmxvYWQoaGVyZTo6aGVyZSgiZGF0YV9maWxlcy9jb3ZfZm9yX21lYW5fdG9fcGxvdF9hbmRfY292X2Zvcl9tZWFuLlJEYXRhIikpCnBsb3QoWV9tZWFuLCB0eXBlID0gImwiLCBjb2wgPSAiZGFya2JsdWUiKQpsaW5lcyhjb3ZfZm9yX21lYW4sIGNvbCA9ICJkYXJrcmVkIikKYGBgCgpCZWxvdyB3ZSBwbG90IHRoZSByZXN1bHRzIGFzIHdlIGRpZCBiZWZvcmUuCgoKCmBgYHtyfQojIFBsb3QgdGhlIGZpZWxkIG9uIHRvcCBvZiB0aGUgbWFwCmYxMiA8LSBncmFwaCRwbG90X2Z1bmN0aW9uKFggPSBjb3ZfZm9yX21lYW5fdG9fcGxvdCwgCiAgICAgICAgICAgICAgICAgICAgICAgICAgdmVydGV4X3NpemUgPSAwLCAKICAgICAgICAgICAgICAgICAgICAgICAgICBwID0gcDEyLAogICAgICAgICAgICAgICAgICAgICAgICAgIGVkZ2Vfd2lkdGggPSAwLjUpICsgCiAgdGhlbWVfbWluaW1hbCgpICsgCiAgdGhlbWUodGV4dCA9IGVsZW1lbnRfdGV4dChmYW1pbHkgPSAiUGFsYXRpbm8iKSwgCiAgICAgICAgYXhpcy50ZXh0ID0gZWxlbWVudF90ZXh0KHNpemUgPSA4KSwKICAgICAgICBsZWdlbmQudGV4dCA9IGVsZW1lbnRfdGV4dChzaXplID0gOCksCiAgICAgICAgcGxvdC5tYXJnaW4gPSB1bml0KC0wLjQqYygxLDAsMSwxKSwgImNtIikKICAgICAgICApICsKICBsYWJzKGNvbG9yID0gIiIsIHggPSAiIiwgeSA9ICIiKSArCiAgeGxpbSh4X2xlZnQsIHhfcmlnaHQpICsgCiAgeWxpbSh5X2JvdHRvbSwgeV90b3ApCiMgUGxvdCB0aGUgZmllbGQgb24gdG9wIG9mIHRoZSBtYXAKZjEzIDwtIGdyYXBoJHBsb3RfZnVuY3Rpb24oWCA9IGNvdl9mb3JfbWVhbl90b19wbG90LCAKICAgICAgICAgICAgICAgICAgICAgICAgICB2ZXJ0ZXhfc2l6ZSA9IDAsIAogICAgICAgICAgICAgICAgICAgICAgICAgIHAgPSBwMTMsCiAgICAgICAgICAgICAgICAgICAgICAgICAgZWRnZV93aWR0aCA9IDAuNSkgKyAKICB0aGVtZV9taW5pbWFsKCkgKyAKICB0aGVtZSh0ZXh0ID0gZWxlbWVudF90ZXh0KGZhbWlseSA9ICJQYWxhdGlubyIpLCAKICAgICAgICBheGlzLnRleHQgPSBlbGVtZW50X3RleHQoc2l6ZSA9IDgpLAogICAgICAgIGxlZ2VuZC50ZXh0ID0gZWxlbWVudF90ZXh0KHNpemUgPSA4KSwKICAgICAgICBwbG90Lm1hcmdpbiA9IHVuaXQoLTAuNCpjKDEsMCwxLDEpLCAiY20iKQogICAgICAgICkgKwogIGxhYnMoY29sb3IgPSAiIiwgeCA9ICIiLCB5ID0gIiIpICsKICB4bGltKHhfbGVmdCwgeF9yaWdodCkgKyAKICB5bGltKHlfYm90dG9tLCB5X3RvcCkKCmcxMiA8LSBncmFwaCRwbG90KGRhdGEgPSAieSIsIHZlcnRleF9zaXplID0gMCwgcCA9IGYxMiwgZWRnZV93aWR0aCA9IDAsIGRhdGFfc2l6ZSA9IDEpICsgCiAgbGFicyhjb2xvciA9ICIiLCB4ID0gIiIsIHkgPSAiIikgKyAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMSwgeSA9IGNvb3JkeV9sd3IxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfbHdyMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIEJvdHRvbSBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjEsIHkgPSBjb29yZHlfdXByMSwgeGVuZCA9IGNvb3JkeF91cHIxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBUb3AgbGluZQogIGFubm90YXRlKCJzZWdtZW50IiwgeCA9IGNvb3JkeF9sd3IxLCB5ID0gY29vcmR5X2x3cjEsIHhlbmQgPSBjb29yZHhfbHdyMSwgeWVuZCA9IGNvb3JkeV91cHIxLCAKICAgICAgICAgICBsaW5ld2lkdGggPSAwLjQsIGNvbG9yID0gdXBwZXJfY29sb3IpICsgICMgTGVmdCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X3VwcjEsIHkgPSBjb29yZHlfbHdyMSwgeGVuZCA9IGNvb3JkeF91cHIxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBSaWdodCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X2x3cjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfdXByMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF9sd3IyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X3VwcjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikKZzEzIDwtIGdyYXBoJHBsb3QoZGF0YSA9ICJ5IiwgdmVydGV4X3NpemUgPSAwLCBwID0gZjEzLCBlZGdlX3dpZHRoID0gMCwgZGF0YV9zaXplID0gMSkgKyAKICBsYWJzKGNvbG9yID0gIiIsIHggPSAiIiwgeSA9ICIiKSArIAogIGFubm90YXRlKCJzZWdtZW50IiwgeCA9IGNvb3JkeF9sd3IxLCB5ID0gY29vcmR5X2x3cjEsIHhlbmQgPSBjb29yZHhfdXByMSwgeWVuZCA9IGNvb3JkeV9sd3IxLCAKICAgICAgICAgICBsaW5ld2lkdGggPSAwLjQsIGNvbG9yID0gdXBwZXJfY29sb3IpICsgICMgQm90dG9tIGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMSwgeSA9IGNvb3JkeV91cHIxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfdXByMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIFRvcCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjEsIHkgPSBjb29yZHlfbHdyMSwgeGVuZCA9IGNvb3JkeF9sd3IxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBMZWZ0IGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfdXByMSwgeSA9IGNvb3JkeV9sd3IxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfdXByMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIFJpZ2h0IGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfbHdyMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV91cHIyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X2x3cjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfdXByMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSAKCnIxIDwtIGcxMyArIHhsaW0oY29vcmR4X2x3cjEsIGNvb3JkeF91cHIxKSArIAogICAgICAgICAgICAgICAgICAgIHlsaW0oY29vcmR5X2x3cjEsIGNvb3JkeV91cHIxKSArIAogIHRoZW1lKGxlZ2VuZC5wb3NpdGlvbiA9ICJub25lIiwgCiAgICAgICAgcGxvdC5tYXJnaW4gPSB1bml0KC0wLjIqYygxLDEsMSwxKSwgImNtIikpCgpyMiA8LSBnMTMgKyB4bGltKGNvb3JkeF9sd3IyLCBjb29yZHhfdXByMikgKyAKICAgICAgICAgICAgICAgICAgICB5bGltKGNvb3JkeV9sd3IyLCBjb29yZHlfdXByMikgKyAKICB0aGVtZShsZWdlbmQucG9zaXRpb24gPSAibm9uZSIsIAogICAgICAgIHBsb3QubWFyZ2luID0gdW5pdCgtMC4yKmMoMSwxLDEsMSksICJjbSIpKQoKIyBBcnJhbmdlIHAyIGFuZCBwMyBob3Jpem9udGFsbHkKbGVmdF9jb2wgPC0gcGxvdF9ncmlkKHIyLCByMSwgbGFiZWxzID0gTlVMTCwgbmNvbCA9IDEsIG5yb3cgPSAyLCByZWxfaGVpZ2h0cyA9IGMoMSwxKSkKCiMgQ29tYmluZSB0aGUgdG9wIHJvdyB3aXRoIHAxIGluIGEgZ3JpZApjb21iaW5lZF9wbG90IDwtIHBsb3RfZ3JpZChsZWZ0X2NvbCwgZzEyLCBsYWJlbHMgPSBOVUxMLCBuY29sID0gMiwgcmVsX3dpZHRocyA9IGMoMSwyKSkgCmZpbmFsX3Bsb3QgPC0gYW5ub3RhdGVfZmlndXJlKGNvbWJpbmVkX3Bsb3QsIGxlZnQgPSB0ZXh0R3JvYigiTGF0aXR1ZGUiLCByb3QgPSA5MCwgdmp1c3QgPSAxLCBncCA9IGdwYXIoY2V4ID0gMC44KSksCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJvdHRvbSA9IHRleHRHcm9iKCJMb25naXR1ZGUiLCB2anVzdCA9IC0wLjUsIGdwID0gZ3BhcihjZXggPSAwLjgpKSkKZ2dzYXZlKGhlcmUoImRhdGFfZmlsZXMvbWVhbl9iZXR0ZXJfMy5wbmciKSwgd2lkdGggPSAxMS4yLCBoZWlnaHQgPSA1LjQzLCBwbG90ID0gZmluYWxfcGxvdCwgZHBpID0gNTAwKQpgYGAKCgpgYGB7ciwgZXZhbCA9IFRSVUUsIGZpZy53aWR0aCA9IDExLjIsIGZpZy5oZWlnaHQgPSA1LjQzLCBmaWcuY2FwID0gY2FwdGlvbmVyKCJBdmVyYWdlIHNwZWVkcyBvZiB0aGUgZmlyc3QgcGFydCBvZiB0aGUgZGF0YS4gUG9pbnRzIHJlcHJlc2VudCB0aGUgcG9pbnQgdmFsdWUgYXQgZWFjaCBhdmFpbGFibGUgbG9jYXRpb24sIGFuZCBlZGdlcyByZXByZXNlbnQgdGhlICRcXG1hdGhybXttZWFuLmNvdn0ocykkIGNvdmFyaWF0ZS4iKX0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MoaGVyZTo6aGVyZSgiZGF0YV9maWxlcy9tZWFuX2JldHRlcl8zLnBuZyIpKQpgYGAKCldlIGZpbmFsbHkgYWRkIHRoZSByZW1haW5pbmcgaGFsZiBvZiB0aGUgcmVwbGljYXRlcyB0byB0aGUgZ3JhcGggYW5kIHBsb3Qgb25lIG9mIHRoZW0uCgpgYGB7cn0KZGZfcmVwIDwtIGxhcHBseSgxOm5yb3coWSksIGZ1bmN0aW9uKGkpe2RhdGEuZnJhbWUoeSA9IFlbaSxdLAogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBtZWFuX3ZhbHVlID0gY292X2Zvcl9tZWFuLAogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZGdlX251bWJlciA9IFB0RVssMV0sCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGRpc3RhbmNlX29uX2VkZ2UgPSBQdEVbLDJdLAogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICByZXBsID0gaSl9KQpkZl9yZXAgPC0gZG8uY2FsbChyYmluZCwgZGZfcmVwKQoKZ3JhcGgkYWRkX29ic2VydmF0aW9ucyhkYXRhID0gZGZfcmVwLCAKICAgICAgICAgICAgICAgICAgICAgICBlZGdlX251bWJlciA9ICJlZGdlX251bWJlciIsCiAgICAgICAgICAgICAgICAgICAgICAgZGlzdGFuY2Vfb25fZWRnZSA9ICJkaXN0YW5jZV9vbl9lZGdlIiwKICAgICAgICAgICAgICAgICAgICAgICBkYXRhX2Nvb3JkcyA9ICJQdEUiLAogICAgICAgICAgICAgICAgICAgICAgIG5vcm1hbGl6ZWQgPSBUUlVFLCAKICAgICAgICAgICAgICAgICAgICAgICBjbGVhcl9vYnMgPSBUUlVFLCAKICAgICAgICAgICAgICAgICAgICAgICBncm91cCA9ICJyZXBsIikKYGBgCgpCZWxvdyB3ZSBwbG90IHJlcGxpY2F0ZSAxNC4KCgoKYGBge3J9CmcxMiA8LSBncmFwaCRwbG90KGRhdGEgPSAieSIsIAogICAgICAgICAgICAgICAgIHAgPSBwMTIsCiAgICAgICAgICAgICAgICAgZ3JvdXAgPSAxLCAKICAgICAgICAgICAgICAgICB2ZXJ0ZXhfc2l6ZSA9IDAsIAogICAgICAgICAgICAgICAgIGRhdGFfc2l6ZSA9IDEpICsgCiAgdGhlbWVfbWluaW1hbCgpICsgCiAgdGhlbWUodGV4dCA9IGVsZW1lbnRfdGV4dChmYW1pbHkgPSAiUGFsYXRpbm8iKSwgCiAgICAgICAgYXhpcy50ZXh0ID0gZWxlbWVudF90ZXh0KHNpemUgPSA4KSwKICAgICAgICBsZWdlbmQudGV4dCA9IGVsZW1lbnRfdGV4dChzaXplID0gOCksCiAgICAgICAgcGxvdC5tYXJnaW4gPSB1bml0KC0wLjQqYygxLDAsMSwxKSwgImNtIikpICsKICBsYWJzKGNvbG9yID0gIiIsIHggPSAiIiwgeSA9ICIiKSArCiAgeGxpbSh4X2xlZnQsIHhfcmlnaHQpICsgCiAgeWxpbSh5X2JvdHRvbSwgeV90b3ApICsKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMSwgeSA9IGNvb3JkeV9sd3IxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfbHdyMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIEJvdHRvbSBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjEsIHkgPSBjb29yZHlfdXByMSwgeGVuZCA9IGNvb3JkeF91cHIxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBUb3AgbGluZQogIGFubm90YXRlKCJzZWdtZW50IiwgeCA9IGNvb3JkeF9sd3IxLCB5ID0gY29vcmR5X2x3cjEsIHhlbmQgPSBjb29yZHhfbHdyMSwgeWVuZCA9IGNvb3JkeV91cHIxLCAKICAgICAgICAgICBsaW5ld2lkdGggPSAwLjQsIGNvbG9yID0gdXBwZXJfY29sb3IpICsgICMgTGVmdCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X3VwcjEsIHkgPSBjb29yZHlfbHdyMSwgeGVuZCA9IGNvb3JkeF91cHIxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBSaWdodCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X2x3cjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfdXByMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF9sd3IyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikgKyAgCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X3VwcjIsIHkgPSBjb29yZHlfbHdyMiwgeGVuZCA9IGNvb3JkeF91cHIyLCB5ZW5kID0gY29vcmR5X3VwcjIsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSBsb3dlcl9jb2xvcikKCmcxMyA8LSBncmFwaCRwbG90KGRhdGEgPSAieSIsIAogICAgICAgICAgICAgICAgIHAgPSBwMTMsCiAgICAgICAgICAgICAgICAgZ3JvdXAgPSAxLCAKICAgICAgICAgICAgICAgICB2ZXJ0ZXhfc2l6ZSA9IDAsIAogICAgICAgICAgICAgICAgIGRhdGFfc2l6ZSA9IDEpICsgCiAgdGhlbWVfbWluaW1hbCgpICsgCiAgdGhlbWUodGV4dCA9IGVsZW1lbnRfdGV4dChmYW1pbHkgPSAiUGFsYXRpbm8iKSwgCiAgICAgICAgYXhpcy50ZXh0ID0gZWxlbWVudF90ZXh0KHNpemUgPSA4KSwKICAgICAgICBsZWdlbmQudGV4dCA9IGVsZW1lbnRfdGV4dChzaXplID0gOCksCiAgICAgICAgcGxvdC5tYXJnaW4gPSB1bml0KC0wLjQqYygxLDAsMSwxKSwgImNtIikpICsKICBsYWJzKGNvbG9yID0gInNwZWVkIiwgeCA9ICIiLCB5ID0gIiIpICsKICB4bGltKHhfbGVmdCwgeF9yaWdodCkgKyAKICB5bGltKHlfYm90dG9tLCB5X3RvcCkgKwogIGFubm90YXRlKCJzZWdtZW50IiwgeCA9IGNvb3JkeF9sd3IxLCB5ID0gY29vcmR5X2x3cjEsIHhlbmQgPSBjb29yZHhfdXByMSwgeWVuZCA9IGNvb3JkeV9sd3IxLCAKICAgICAgICAgICBsaW5ld2lkdGggPSAwLjQsIGNvbG9yID0gdXBwZXJfY29sb3IpICsgICMgQm90dG9tIGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMSwgeSA9IGNvb3JkeV91cHIxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfdXByMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIFRvcCBsaW5lCiAgYW5ub3RhdGUoInNlZ21lbnQiLCB4ID0gY29vcmR4X2x3cjEsIHkgPSBjb29yZHlfbHdyMSwgeGVuZCA9IGNvb3JkeF9sd3IxLCB5ZW5kID0gY29vcmR5X3VwcjEsIAogICAgICAgICAgIGxpbmV3aWR0aCA9IDAuNCwgY29sb3IgPSB1cHBlcl9jb2xvcikgKyAgIyBMZWZ0IGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfdXByMSwgeSA9IGNvb3JkeV9sd3IxLCB4ZW5kID0gY29vcmR4X3VwcjEsIHllbmQgPSBjb29yZHlfdXByMSwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IHVwcGVyX2NvbG9yKSArICAjIFJpZ2h0IGxpbmUKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfbHdyMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV91cHIyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfbHdyMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X2x3cjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKSArICAKICBhbm5vdGF0ZSgic2VnbWVudCIsIHggPSBjb29yZHhfdXByMiwgeSA9IGNvb3JkeV9sd3IyLCB4ZW5kID0gY29vcmR4X3VwcjIsIHllbmQgPSBjb29yZHlfdXByMiwgCiAgICAgICAgICAgbGluZXdpZHRoID0gMC40LCBjb2xvciA9IGxvd2VyX2NvbG9yKQoKcjEgPC0gZzEzICsgeGxpbShjb29yZHhfbHdyMSwgY29vcmR4X3VwcjEpICsgCiAgICAgICAgICAgICAgICAgICAgeWxpbShjb29yZHlfbHdyMSwgY29vcmR5X3VwcjEpICsgCiAgdGhlbWUobGVnZW5kLnBvc2l0aW9uID0gIm5vbmUiLCAKICAgICAgICBwbG90Lm1hcmdpbiA9IHVuaXQoLTAuMipjKDEsMSwxLDEpLCAiY20iKSkKCnIyIDwtIGcxMyArIHhsaW0oY29vcmR4X2x3cjIsIGNvb3JkeF91cHIyKSArIAogICAgICAgICAgICAgICAgICAgIHlsaW0oY29vcmR5X2x3cjIsIGNvb3JkeV91cHIyKSArIAogIHRoZW1lKGxlZ2VuZC5wb3NpdGlvbiA9ICJub25lIiwgCiAgICAgICAgcGxvdC5tYXJnaW4gPSB1bml0KC0wLjIqYygxLDEsMSwxKSwgImNtIikpCgojIEFycmFuZ2UgcDIgYW5kIHAzIGhvcml6b250YWxseQpsZWZ0X2NvbCA8LSBwbG90X2dyaWQocjIsIHIxLCBsYWJlbHMgPSBOVUxMLCBuY29sID0gMSwgbnJvdyA9IDIsIHJlbF9oZWlnaHRzID0gYygxLDEpKQoKIyBDb21iaW5lIHRoZSB0b3Agcm93IHdpdGggcDEgaW4gYSBncmlkCmNvbWJpbmVkX3Bsb3QgPC0gcGxvdF9ncmlkKGxlZnRfY29sLCBnMTIsIGxhYmVscyA9IE5VTEwsIG5jb2wgPSAyLCByZWxfd2lkdGhzID0gYygxLDIpKSAKZmluYWxfcGxvdCA8LSBhbm5vdGF0ZV9maWd1cmUoY29tYmluZWRfcGxvdCwgbGVmdCA9IHRleHRHcm9iKCJMYXRpdHVkZSIsIHJvdCA9IDkwLCB2anVzdCA9IDEsIGdwID0gZ3BhcihjZXggPSAwLjgpKSwKICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYm90dG9tID0gdGV4dEdyb2IoIkxvbmdpdHVkZSIsIHZqdXN0ID0gLTAuNSwgZ3AgPSBncGFyKGNleCA9IDAuOCkpKQpnZ3NhdmUoaGVyZSgiZGF0YV9maWxlcy9yZXBsaWNhdGUxNF8zLnBuZyIpLCB3aWR0aCA9IDExLjIsIGhlaWdodCA9IDUuNDMsIHBsb3QgPSBmaW5hbF9wbG90LCBkcGkgPSA1MDApCmBgYAoKCmBgYHtyLCBldmFsID0gVFJVRSwgZmlnLndpZHRoID0gMTEuMiwgZmlnLmhlaWdodCA9IDUuNDMsIGZpZy5jYXAgPSBjYXB0aW9uZXIoIlNwZWVkIG9ic2VydmF0aW9ucyAoaW4gbXBoKSBvbiB0aGUgaGlnaHdheSBuZXR3b3JrIG9mIHRoZSBjaXR5IG9mIFNhbiBKb3NlIGluIENhbGlmb3JuaWEsIHJlY29yZGVkIG9uIEFwcmlsIDMsIDIwMTcuIFRoZSBsZWZ0IHBhbmVscyBhcmUgem9vbWVkLWluIGFyZWFzIG9mIHRoZSBwYW5lbCB0byB0aGUgcmlnaHQuIil9CmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKGhlcmU6OmhlcmUoImRhdGFfZmlsZXMvcmVwbGljYXRlMTRfMy5wbmciKSkKYGBgCgoKYGBge3J9CnNhdmUoZ3JhcGgsIGNvdiwgY292X2Zvcl9tZWFuX3RvX3Bsb3QsIGRhdGFfcHJkX2xpc3RfbWVzaCwgZmlsZSA9IGhlcmUoImRhdGFfZmlsZXMvcGVtc19yZXBsMV9kYXRhLlJEYXRhIikpCmBgYAoKCiMgUmVmZXJlbmNlcwoKCmBgYHtyLCBldmFsID0gVFJVRX0KZ3JhdGVmdWw6OmNpdGVfcGFja2FnZXMob3V0cHV0ID0gInBhcmFncmFwaCIsIG91dC5kaXIgPSAiLiIpCmBgYAoKCg==